Fontenot, J. D., Gavin, M. A. & Rudensky, A. Y. FoxP3 programs the development and function of CD4+CD25+ regulatory T cells. Nat. Immunol. 4, 330–336 (2003).
Hori, S., Nomura, T. & Sakaguchi, S. Control of regulatory T cell development by the transcription factor Foxp3. Science 299, 1057–1061 (2003).
Khattri, R., Cox, T., Yasayko, S. A. & Ramsdell, F. An essential role for Scurfin in CD4+CD25+ T regulatory cells. Nat. Immunol. 4, 337–342 (2003).
Brunkow, M. E. et al. Disruption of a new forkhead/winged-helix protein, scurfin, results in the fatal lymphoproliferative disorder of the scurfy mouse. Nat. Genet. 27, 68–73 (2001).
Chatila, T. A. et al. JM2, encoding a fork head-related protein, is mutated in X-linked autoimmunity-allergic disregulation syndrome. J. Clin. Invest. 106, R75–R81 (2000).
Bennett, C. L. et al. The immune dysregulation, polyendocrinopathy, enteropathy, X-linked syndrome (IPEX) is caused by mutations of FOXP3. Nat. Genet. 27, 20–21 (2001).
Wildin, R. S. et al. X-linked neonatal diabetes mellitus, enteropathy and endocrinopathy syndrome is the human equivalent of mouse scurfy. Nat. Genet. 27, 18–20 (2001).
Bacchetta, R. & Roncarolo, M. G. IPEX syndrome from diagnosis to cure, learning along the way. J. Allergy Clin. Immunol. 153, 595–605 (2024).
Lin, W. et al. Regulatory T cell development in the absence of functional Foxp3. Nat. Immunol. 8, 359–368 (2007).
Gavin, M. A. et al. Foxp3-dependent programme of regulatory T-cell differentiation. Nature 445, 771–775 (2007).
Hill, J. A. et al. Foxp3 transcription-factor-dependent and -independent regulation of the regulatory T cell transcriptional signature. Immunity 27, 786–800 (2007).
Hill, J. A. et al. Retinoic acid enhances Foxp3 induction indirectly by relieving inhibition from CD4+CD44hi cells. Immunity 29, 758–770 (2008).
van der Veeken, J. et al. The transcription factor Foxp3 shapes regulatory T cell identity by tuning the activity of trans-acting intermediaries. Immunity 53, 971–984 (2020).
Kwon, H.-K., Chen, H.-M., Mathis, D. & Benoist, C. Different molecular complexes that mediate transcriptional induction and repression by FoxP3. Nat. Immunol. 18, 1238–1248 (2017).
Marson, A. et al. Foxp3 occupancy and regulation of key target genes during T-cell stimulation. Nature 445, 931–935 (2007).
Zheng, Y. et al. Genome-wide analysis of Foxp3 target genes in developing and mature regulatory T cells. Nature 445, 936–940 (2007).
Yoshida, H. et al. The cis-regulatory atlas of the mouse immune system. Cell 176, 897–912 (2019).
Zemmour, D. et al. Single-cell analysis of FOXP3 deficiencies in humans and mice unmasks intrinsic and extrinsic CD4+ T cell perturbations. Nat. Immunol. 22, 607–619 (2021).
Hu, W. et al. Temporal and context-dependent requirements for the transcription factor Foxp3 expression in regulatory T cells. Nat. Immunol. 26, 2059–2073 (2025).
Jager, C. Inducible protein degradation reveals inflammation-dependent function of the Treg cell lineage-defining transcription factor Foxp3. Sci. Immunol. 10, eadr7057 (2025).
Arvey, A. et al. Inflammation-induced repression of chromatin bound by the transcription factor Foxp3 in regulatory T cells. Nat. Immunol. 15, 580–587 (2014).
Peeters, J. G. C., Silveria, S., Ozdemir, M., Ramachandran, S. & DuPage, M. Hyperactivating EZH2 to augment H3K27me3 levels in regulatory T cells enhances immune suppression by driving early effector differentiation. Cell Rep. 43, 114724 (2024).
DuPage, M. et al. The chromatin-modifying enzyme EZH2 is critical for the maintenance of regulatory T cell identity after activation. Immunity 42, 227–238 (2015).
Yang, X. P. et al. EZH2 is crucial for both differentiation of regulatory T cells and T effector cell expansion. Sci. Rep. 5, 10643 (2015).
Sarmento, O. F. et al. The role of the histone methyltransferase enhancer of zeste homolog 2 (EZH2) in the pathobiological mechanisms underlying inflammatory bowel disease (IBD). J. Biol. Chem. 292, 706–722 (2017).
Zhang, Y. et al. The polycomb repressive complex 2 governs life and death of peripheral T cells. Blood 124, 737–749 (2014).
Wang, D. et al. Targeting EZH2 reprograms intratumoral regulatory T cells to enhance cancer immunity. Cell Rep. 23, 3262–3274 (2018).
Samstein, R. M. et al. Foxp3 exploits a pre-existent enhancer landscape for regulatory T cell lineage specification. Cell 151, 153–166 (2012).
Liu, Z., Lee, D. S., Liang, Y., Zheng, Y. & Dixon, J. R. Foxp3 orchestrates reorganization of chromatin architecture to establish regulatory T cell identity. Nat. Commun. 14, 6943 (2023).
Kitagawa, Y. et al. Guidance of regulatory T cell development by SATB1-dependent super-enhancer establishment. Nat. Immunol. 18, 173–183 (2017).
Arora, S. et al. Joint sequence and chromatin neural networks characterize the differential abilities of forkhead transcription factors to engage inaccessible chromatin. Preprint at bioRxiv https://doi.org/10.1101/2023.10.06.561228 (2023).
DelRosso, N. et al. Large-scale mapping and mutagenesis of human transcriptional effector domains. Nature 616, 365–372 (2023).
Du, J., Huang, C., Zhou, B. & Ziegler, S. F. Isoform-specific inhibition of RORα-mediated transcriptional activation by human FOXP3. J. Immunol. 180, 4785–4792 (2008).
Ichiyama, K. et al. Foxp3 inhibits RORγt-mediated IL-17A mRNA transcription through direct interaction with RORγt. J. Biol. Chem. 283, 17003–17008 (2008).
Huang, C. et al. Cutting edge: A novel, human-specific interacting protein couples FOXP3 to a chromatin-remodeling complex that contains KAP1/TRIM28. J. Immunol. 190, 4470–4473 (2013).
Li, B. et al. FOXP3 interactions with histone acetyltransferase and class II histone deacetylases are required for repression. Proc. Natl Acad. Sci. USA 104, 4571–4576 (2007).
Pan, F. et al. Eos mediates Foxp3-dependent gene silencing in CD4+ regulatory T cells. Science 325, 1142–1146 (2009).
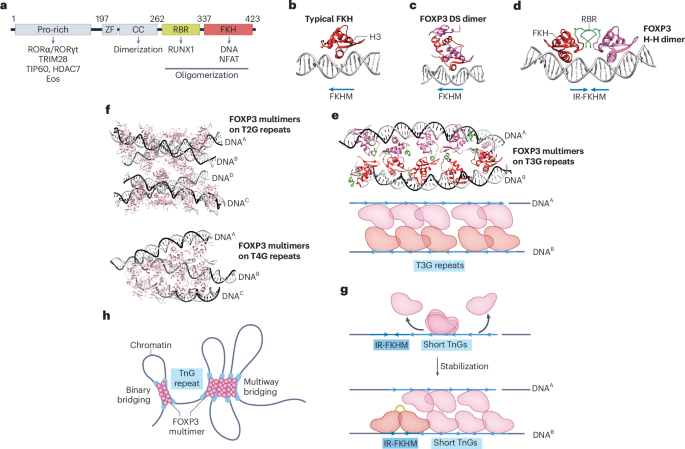
Leng, F. et al. The transcription factor FoxP3 can fold into two dimerization states with divergent implications for regulatory T cell function and immune homeostasis. Immunity 55, 1354–1369 (2022).
Song, X. et al. Structural and biological features of FOXP3 dimerization relevant to regulatory T cell function. Cell Rep. 1, 665–675 (2012).
Ono, M. et al. Foxp3 controls regulatory T-cell function by interacting with AML1/Runx1. Nature 446, 685–689 (2007).
Rudra, D. et al. Runx–CBFβ complexes control expression of the transcription factor Foxp3 in regulatory T cells. Nat. Immunol. 10, 1170–1177 (2009).
Zhang, W. et al. FOXP3 recognizes microsatellites and bridges DNA through multimerization. Nature 624, 433–441 (2023).
Leng, F. et al. Ultrastable and versatile multimeric ensembles of FoxP3 on microsatellites. Mol. Cell 85, 1509–1524 (2025).
Dai, S., Qu, L., Li, J. & Chen, Y. Toward a mechanistic understanding of DNA binding by forkhead transcription factors and its perturbation by pathogenic mutations. Nucleic Acids Res. 49, 10235–10249 (2021).
Bandukwala, H. S. et al. Structure of a domain-swapped FOXP3 dimer on DNA and its function in regulatory T cells. Immunity 34, 479–491 (2011).
Chen, Y. et al. DNA binding by FOXP3 domain-swapped dimer suggests mechanisms of long-range chromosomal interactions. Nucleic Acids Res. 43, 1268–1282 (2015).
Wu, Y. et al. FOXP3 controls regulatory T cell function through cooperation with NFAT. Cell 126, 375–387 (2006).
Chu, Y. P. et al. Solution structure and backbone dynamics of the DNA-binding domain of FOXP1: insight into its domain swapping and DNA binding. Protein Sci. 20, 908–924 (2011).
Stroud, J. C. et al. Structure of the forkhead domain of FOXP2 bound to DNA. Structure 14, 159–166 (2006).
Van Gool, F. et al. A mutation in the transcription factor Foxp3 drives T helper 2 effector function in regulatory T cells. Immunity 50, 362–377 (2019).
Leng, F. et al. FoxP3 forms a head-to-head dimer in vivo and stabilizes its multimerization on adjacent microsatellites. Cell Rep. 44, 116633 (2025).
Nakagawa, S., Gisselbrecht, S. S., Rogers, J. M., Hartl, D. L. & Bulyk, M. L. DNA-binding specificity changes in the evolution of forkhead transcription factors. Proc. Natl Acad. Sci. USA 110, 12349–12354 (2013).
Ramirez, R. N., Chowdhary, K., Leon, J., Mathis, D. & Benoist, C. FoxP3 associates with enhancer–promoter loops to regulate Treg-specific gene expression. Sci. Immunol. 7, eabj9836 (2022).
Nora, E. P. et al. Targeted degradation of CTCF decouples local insulation of chromosome domains from genomic compartmentalization. Cell 169, 930–944 (2017).
Ghavi-Helm, Y. et al. Highly rearranged chromosomes reveal uncoupling between genome topology and gene expression. Nat. Genet. 51, 1272–1282 (2019).
Rao, S. S. P. et al. Cohesin loss eliminates all loop domains. Cell 171, 305–320 (2017).
Horton, C. A. et al. Short tandem repeats bind transcription factors to tune eukaryotic gene expression. Science 381, eadd1250 (2023).
Xing, Y. & Hogquist, K. A. T-cell tolerance: central and peripheral. Cold Spring Harb. Perspect. Biol. 4, a006957 (2012).
Derbinski, J., Schulte, A., Kyewski, B. & Klein, L. Promiscuous gene expression in medullary thymic epithelial cells mirrors the peripheral self. Nat. Immunol. 2, 1032–1039 (2001).
Sansom, S. N. et al. Population and single-cell genomics reveal the Aire dependency, relief from Polycomb silencing, and distribution of self-antigen expression in thymic epithelia. Genome Res. 24, 1918–1931 (2014).
Anderson, M. S. et al. Projection of an immunological self shadow within the thymus by the Aire protein. Science 298, 1395–1401 (2002).
Malchow, S. et al. Aire enforces immune tolerance by directing autoreactive T cells into the regulatory T cell lineage. Immunity 44, 1102–1113 (2016).
Yang, S., Fujikado, N., Kolodin, D., Benoist, C. & Mathis, D. Regulatory T cells generated early in life play a distinct role in maintaining self-tolerance. Science 348, 589–594 (2015).
Brennecke, P. et al. Single-cell transcriptome analysis reveals coordinated ectopic gene-expression patterns in medullary thymic epithelial cells. Nat. Immunol. 16, 933–941 (2015).
Meredith, M., Zemmour, D., Mathis, D. & Benoist, C. Aire controls gene expression in the thymic epithelium with ordered stochasticity. Nat. Immunol. 16, 942–949 (2015).
Bautista, J. L. et al. Single-cell transcriptional profiling of human thymic stroma uncovers novel cellular heterogeneity in the thymic medulla. Nat. Commun. 12, 1096 (2021).
Bornstein, C. et al. Single-cell mapping of the thymic stroma identifies IL-25-producing tuft epithelial cells. Nature 559, 622–626 (2018).
Givony, T. et al. Thymic mimetic cells function beyond self-tolerance. Nature 622, 164–172 (2023).
Michelson, D. A., Hase, K., Kaisho, T., Benoist, C. & Mathis, D. Thymic epithelial cells co-opt lineage-defining transcription factors to eliminate autoreactive T cells. Cell 185, 2542–2558 (2022).
Miller, C. N. et al. Thymic tuft cells promote an IL-4-enriched medulla and shape thymocyte development. Nature 559, 627–631 (2018).
Michelson, D. A., Zuo, C., Verzi, M., Benoist, C. & Mathis, D. HNF4 activates mimetic-cell enhancers to recapitulate gut and liver development within the thymus. J. Exp. Med. 220, e20230461 (2023).
Sin, J. H. et al. Ikaros is a principal regulator of Aire+ mTEC homeostasis, thymic mimetic cell diversity, and central tolerance. Sci. Immunol. 8, eabq3109 (2023).
Husebye, E. S., Anderson, M. S. & Kampe, O. Autoimmune polyendocrine syndromes. N. Engl. J. Med. 378, 1132–1141 (2018).
Metzger, T. C. et al. Lineage tracing and cell ablation identify a post-Aire-expressing thymic epithelial cell population. Cell Rep. 5, 166–179 (2013).
Abramson, J., Giraud, M., Benoist, C. & Mathis, D. Aire’s partners in the molecular control of immunological tolerance. Cell 140, 123–135 (2010).
Berger, A. H. et al. High-resolution transcriptional impact of AIRE: effects of pathogenic variants p.Arg257Ter, p.Cys311Tyr, and polygenic risk variant p.Arg471Cys. Front. Immunol. 16, 1572789 (2025).
Giraud, M. et al. Aire unleashes stalled RNA polymerase to induce ectopic gene expression in thymic epithelial cells. Proc. Natl Acad. Sci. USA 109, 535–540 (2012).
Huoh, Y. S. et al. Dual functions of Aire CARD multimerization in the transcriptional regulation of T cell tolerance. Nat. Commun. 11, 1625 (2020).
Huoh, Y. S. et al. Mechanism for controlled assembly of transcriptional condensates by Aire. Nat. Immunol. 25, 1580–1592 (2024).
Koh, A. S. et al. Aire employs a histone-binding module to mediate immunological tolerance, linking chromatin regulation with organ-specific autoimmunity. Proc. Natl Acad. Sci. USA 105, 15878–15883 (2008).
Org, T. et al. The autoimmune regulator PHD finger binds to non-methylated histone H3K4 to activate gene expression. EMBO Rep. 9, 370–376 (2008).
Lim, S. et al. Derivation of functional thymic epithelial organoid lines from adult murine thymus. Cell Rep. 43, 114019 (2024).
Hubscher, T. et al. Thymic epithelial organoids mediate T-cell development. Development 151, dev202853 (2024).
Major, V. et al. Establishment of a microwell-array-based miniaturized thymic organoid model suitable for high-throughput applications. Cell Rep. 44, 115579 (2025).
Ramos, S. A. et al. Generation of functional thymic organoids from human pluripotent stem cells. Stem Cell Reports 18, 829–840 (2023).
Waterfield, M. et al. The transcriptional regulator Aire coopts the repressive ATF7ip-MBD1 complex for the induction of immunotolerance. Nat. Immunol. 15, 258–265 (2014).
Meloni, A., Incani, F., Corda, D., Cao, A. & Rosatelli, M. C. Role of PHD fingers and COOH-terminal 30 amino acids in AIRE transactivation activity. Mol. Immunol. 45, 805–809 (2008).
Bruserud, O., Oftedal, B. E., Wolff, A. B. & Husebye, E. S. AIRE—mutations and autoimmune disease. Curr. Opin. Immunol. 43, 8–15 (2016).
Oftedal, B. E. et al. Dominant mutations in the autoimmune regulator AIRE are associated with common organ-specific autoimmune diseases. Immunity 42, 1185–1196 (2015).
Abbott, J. K. et al. Dominant-negative loss of function arises from a second, more frequent variant within the SAND domain of autoimmune regulator (AIRE). J. Autoimmun. 88, 114–120 (2018).
Ochoa, S. et al. A deep intronic splice-altering AIRE variant causes APECED syndrome through antisense oligonucleotide-targetable pseudoexon inclusion. Sci. Transl. Med. 16, eadk0845 (2024).
Meloni, A. et al. Delineation of the molecular defects in the AIRE gene in autoimmune polyendocrinopathy-candidiasis-ectodermal dystrophy patients from Southern Italy. J. Clin. Endocrinol. Metab. 87, 841–846 (2002).
Sabari, B. R., Dall’Agnese, A. & Young, R. A. Biomolecular condensates in the nucleus. Trends Biochem. Sci. 45, 961–977 (2020).
Bjorses, P. et al. Localization of the APECED protein in distinct nuclear structures. Hum. Mol. Genet. 8, 259–266 (1999).
Rinderle, C., Christensen, H. M., Schweiger, S., Lehrach, H. & Yaspo, M. L. AIRE encodes a nuclear protein co-localizing with cytoskeletal filaments: altered sub-cellular distribution of mutants lacking the PHD zinc fingers. Hum. Mol. Genet. 8, 277–290 (1999).
Su, M. A. et al. Mechanisms of an autoimmunity syndrome in mice caused by a dominant mutation in Aire. J. Clin. Invest. 118, 1712–1726 (2008).
Pitkanen, J. et al. Cooperative activation of transcription by autoimmune regulator AIRE and CBP. Biochem. Biophys. Res. Commun. 333, 944–953 (2005).
Yoshida, H. et al. Brd4 bridges the transcriptional regulators, Aire and P-TEFb, to promote elongation of peripheral-tissue antigen transcripts in thymic stromal cells. Proc. Natl Acad. Sci. USA 112, E4448–E4457 (2015).
Bansal, K., Yoshida, H., Benoist, C. & Mathis, D. The transcriptional regulator Aire binds to and activates super-enhancers. Nat. Immunol. 18, 263–273 (2017).
Noma, K., Allis, C. D. & Grewal, S. I. Transitions in distinct histone H3 methylation patterns at the heterochromatin domain boundaries. Science 293, 1150–1155 (2001).
Rea, S. et al. Regulation of chromatin structure by site-specific histone H3 methyltransferases. Nature 406, 593–599 (2000).
Saad, S. et al. DNA binding and mitotic phosphorylation protect polyglutamine proteins from assembly formation. Cell 188, 2974–2991 (2025).
Oftedal, B. E. et al. Dominant-negative heterozygous mutations in AIRE confer diverse autoimmune phenotypes. iScience 26, 106818 (2023).
Koh, A. S., Kingston, R. E., Benoist, C. & Mathis, D. Global relevance of Aire binding to hypomethylated lysine-4 of histone-3. Proc. Natl Acad. Sci. USA 107, 13016–13021 (2010).
Fang, Y., Bansal, K., Mostafavi, S., Benoist, C. & Mathis, D. Aire relies on Z-DNA to flag gene targets for thymic T cell tolerization. Nature 628, 400–407 (2024).
Du, M. et al. Direct observation of a condensate effect on super-enhancer controlled gene bursting. Cell 187, 2595–2598 (2024).
Oven, I. et al. AIRE recruits P-TEFb for transcriptional elongation of target genes in medullary thymic epithelial cells. Mol. Cell. Biol. 27, 8815–8823 (2007).
Takahashi, H. et al. Human mediator subunit MED26 functions as a docking site for transcription elongation factors. Cell 146, 92–104 (2011).
Giraud, M. et al. An RNAi screen for Aire cofactors reveals a role for Hnrnpl in polymerase release and Aire-activated ectopic transcription. Proc. Natl Acad. Sci. USA 111, 1491–1496 (2014).
Keane, P., Ceredig, R. & Seoighe, C. Promiscuous mRNA splicing under the control of AIRE in medullary thymic epithelial cells. Bioinformatics 31, 986–990 (2015).
Handel, A. E. et al. Comprehensively profiling the chromatin architecture of tissue restricted antigen expression in thymic epithelial cells over development. Front. Immunol. 9, 2120 (2018).
Gamble, N. et al. Thymic epithelial cells amplify epigenetic noise to promote immune tolerance. Nature 646, 724–733 (2025).
Michelson, D. A. & Mathis, D. Thymic mimetic cells: ontogeny as immunology. Annu. Rev. Cell Dev. Biol. 40, 283–300 (2024).
Gardner, J. M. et al. Deletional tolerance mediated by extrathymic Aire-expressing cells. Science 321, 843–847 (2008).
Gardner, J. M. et al. Extrathymic Aire-expressing cells are a distinct bone marrow-derived population that induce functional inactivation of CD4+ T cells. Immunity 39, 560–572 (2013).
Akagbosu, B. et al. Novel antigen-presenting cell imparts Treg-dependent tolerance to gut microbiota. Nature 610, 752–760 (2022).
Wang, J. et al. Single-cell multiomics defines tolerogenic extrathymic Aire-expressing populations with unique homology to thymic epithelium. Sci. Immunol. 6, eabl5053 (2021).
Dobes, J. et al. Extrathymic expression of Aire controls the induction of effective TH17 cell-mediated immune response to Candida albicans. Nat. Immunol. 23, 1098–1108 (2022).
Pretemer, Y. et al. An iPSC-based in vitro model recapitulates human thymic epithelial development and multi-lineage specification. Nat. Commun. 16, 7680 (2025).
Fraschilla, I. & Jeffrey, K. L. The speckled protein (SP) family: immunity’s chromatin readers. Trends Immunol. 41, 572–585 (2020).















Leave a Reply