Mice
All mice used in this study were of the C57BL/6 genetic background. Experiments were conducted using age- and sex-matched mice between 6–12 weeks of age. The XKR8 germline KO, floxed and 2DA mutant KI mice were generated for this study. To produce the Xkr8 germline KO mice, exons 2 and 3 of the WT Xkr8 allele were deleted using CRISPR–Cas9 system, as illustrated in Extended Data Fig. 1f. Littermate Xkr8+/+ mice were used as WT controls. To generate neutrophil-specific Xkr8 KO mice (Xkr8fl/flLy6gcre), Xkr8 floxed mice (Xkr8fl/fl) were crossed with Ly6gcre KI mice, as depicted in Extended Data Fig. 9c. The Ly6gcre mice were provided by Z. Zeng. The Xkr82DA (p.D351A&D354A) KI mice were generated via CRISPR–Cas9-mediated KI of the 2DA mutant cassette into exon 3 of Xkr8 gene (see Supplementary Fig. 2a for the detailed strategy). TMEM16F germline KO mice were previously described59. Caspase-3 KO mice and Padi4 KO mice were gifts from S. Shi and X. Wang, respectively. Both male and female mice were used in this study. Mice were housed under specific pathogen-free conditions, and all animal experiments were conducted in accordance with protocols approved by the Animal Care and Use Committee of Anhui Medical University (LLSC20230277).
Antibodies and reagents
Anti-citrullinated histone H3 (citrulline R2 + R8 + R17) (ab5103, 1:1,000 dilution; ab281584, 1:1,000 dilution) and anti-neutrophil elastase (ab314916, 1:100 dilution) were purchased from Abcam. Anti-MPO (AF3667) was obtained from R&D systems. Antibodies against caspase-3 (9662), caspase-7 (D6H1, 8438), caspase-8 (D5B2, 8592), caspase-9 (9504), phospho-ERKT202/Y204 (4370) and phospho-PKC (βII Ser660, 9371) were from Cell Signaling Technologies. The GFP-tag polyclonal antibody (50430-2-AP) and β-actin monoclonal antibody (66009-1-lg) were from Proteintech. Flow cytometry and surface marker antibodies, including those against mouse Ly6G (1A8), CD11a (M17/4), CD11b (M1/70), CD45 (30-F11), CD44 (IM7), CD62L (MEL-14), CD101 (Moushi101), CXCR2 (SA044G4), CXCR4 (2B11), TLR4 (SA15-21), annexin V (640941, 640945) and corresponding isotype controls, were obtained from BioLegend. Elastase G-2 (sc-55549) was obtained from Santa Cruz. Activated caspase-3 antibody (570524), CD16 (555406) and CD66b (561650) were from BD Biosciences.
Chemical compounds
To investigate the signaling pathways involved in NET formation, various chemical inducers and inhibitors were employed. Neutrophils were pretreated with inhibitors for 30 min at 37 °C in serum-free RPMI 1640 (Gibco) supplemented with 10 mM HEPES before stimulation with NET inducers. The following chemical compounds were used in the experiments: PMA (P8139), EGTA (E4378), BAPTA-AM (A1076) and TEMPO (426369) were from Sigma-Aldrich. LPS (tlrl-eblps), nigericin (tlrl-nig), monosodium urate (MSU) crystals (tlrl-msu) and zymosan (tlrl-zyn) were from Invivogen. A23187 (B6646), Q-VD-OPh (A1901), Z-VAD-FMK (A1902), apocynin (B3675), DPI (B6326), U0126 (A1337), GW5074 (B1405), EGTA-AM (C4548), doxorubicin (A1832) and staurosporine (STS, A8192) were all from APExBIO Technology. Additional inhibitors, including S62845 (HY-100741), emricasan (HY-10396), 2-APB (B6643), SKF96365 (HY-100001), HC067047 (HY-100208), Pyr3 (HY-108465), JNJ-28583113 (HY-149143), 9-phenanthrol (HY-108457), carvacrol (HY-N0711), GsMTx-4 (HY-P1410), probenecid (HY-B0545), GSK1016790A (HY-19608), GSK1702934A (HY-111098), bilirubin (HY-N0323), Yoda1 (HY-18723) and NTB (HY-101302) were obtained from MedChemExpress. CBD (M9136) was purchased from Abmole, and the compound NA-11 was previously reported36. Enzastaurin (S1055) and SAR7334 (S6634) were obtained from Selleck, and CM-4620 (CSN21369) and GSK2795039 (33777) were sourced from CSNpharm and Cayman Chemical, respectively. CellEvent Caspase-3/7 Green Detection Reagent (C10423) and Laurdan (D250) were from Invitrogen, and Flipper-TR (CY-SC020) was from Cytoskeleton.
Cell culture and differentiation
HL-60 cells (CCL-240), obtained from ATCC, were cultured at 37 °C with 5% CO2 in IMDM (Gibco) supplemented with 25 mM HEPES, 2 mM l-glutamine, 20% heat-inactivated fetal bovine serum (FBS) and 1% penicillin–streptomycin. Cells were routinely tested negative for Mycoplasma contamination. Differentiation of HL-60 cells into neutrophil-like cells was induced by treating the cells with 10 μM ATRA for 6 days. The efficiency of differentiation was regularly assessed by flow cytometry (CD16+CD66b+). Fully differentiated (dHL-60) cells were used in subsequent functional assays.
Generation of XKR8-deficient HL-60 cells
XKR8-deficient HL-60 cells were generated using CRISPR–Cas9 technology. The LentiCRISPR v2 vector (pLenti-V2, 52961) and packaging plasmids pMD2.G (12259) and psPAX2 (12260) were obtained from Addgene. Gene-specific sgRNAs were designed and cloned into LentiCRISPR v2 vector, with at least two distinct sgRNAs targeting the XKR8 gene. The lentiviral vector and packaging plasmids were co-transfected into HEK293T cells using polyethyleneimine (PEI). After 48 h, viral supernatants were collected, filtered through a 0.22-μm filter and supplemented with Polybrene (5 μg ml−1) for transduction of HL-60 cells. Transduced cells were selected using puromycin (1 μg ml−1). In addition, CRISPR–Cas9 tracrRNA (1072532), Cas9-GFP V3 (10008100) and corresponding CRISPR–Cas9 crRNA were purchased from IDT. The Cas9–GFP protein, tracrRNA and gene-specific crRNAs were co-transfected into HL-60 cells by electroporation (Neon NxT Electroporation System, Thermo Fisher Scientific). Positive cells were sorted by GFP+ (BD AriaⅢ). KO clones were further screened by PCR, and indel mutations in selected clones were verified by genomic DNA sequencing. At least three independent clones were pooled for subsequent experiments. Oligonucleotides used in this study are listed in Supplementary Table 2.
Overexpression of XKR8 and its mutants in HL-60 KO cell lines were achieved using lentivirus-based infection. The cDNA of Xkr8 and its 2DA, S/T-3A, D30A and W45A mutants were cloned into pLVX-IRES-ZsGreen1 (Clontech, Cat. 632187). Transfection and infection processes were performed as previously described. ZsGreen+ cells were sorted for further experiments.
Culture of bacteria and fungi
L. monocytogenes (10403s) was cultured overnight at 37 °C as previously described59. S. aureus (MW2) was cultured overnight at 37 °C in Brain Heart Infusion broth supplemented with erythromycin (10 μg ml−1). To prepare bacteria at the logarithmic phase, 100 μl of the overnight culture was added to 5 ml of fresh medium and incubated for 2 h at 37 °C with agitation. Bacterial cultures were collected when the optical density (OD) at 600 nm reached 0.4, washed with PBS and prepared for subsequent experiments. E. coli (CFT073) was cultured overnight at 37 °C in Luria-Bertani (LB) medium containing kanamycin (50 μg ml−1). For experiments, 100 μl bacterial culture was added to 2 ml of fresh LB medium containing with 1 μM IPTG to boost for 2 h at 37 °C. The bacterial concentration was determined using the formula: 1 OD600nm = 5 × 108 c.f.u. per ml. C. albicans (SC5314) was cultured overnight at 37 °C in yeast extract peptone dextrose (YPD) medium. To boost fungal growth, 100 μl of the overnight fungal culture was transferred to 5 ml of fresh medium and incubated for 2 h at 37 °C. Fungal cells were then collected and counted using a hemocytometer.
Isolation of neutrophils from mouse bone marrow
Neutrophils were isolated from mouse bone marrow using gradient centrifugation with Percoll (17089109-1, GE Healthcare). Bone marrow cells were collected in HBSS supplemented with 0.5% FBS and 20 mM HEPES, without Ca2+ and Mg2+. To remove red blood cells, the pellet was resuspended in 6 ml of 0.2% NaCl for 45 s, followed by the addition of 14 ml of 1.2% NaCl to restore isotonicity. After centrifugation at 1,000g for 30 min at room temperature, the neutrophil-enriched cell pellet, appearing as a cloudy layer at the bottom of the tube, was collected and washed with HBSS. The purity of the neutrophil population was typically greater than 90%, as determined by morphological examination with Giemsa staining and flow cytometry, based on CD11b+ (1:500, clone M1/70) and Ly6G+ (1:500, clone 1A8) surface markers.
Isolation of neutrophils from human peripheral blood
Peripheral blood from healthy donors were obtained from the Department of Clinical Laboratory, the First Affiliated Hospital of Anhui Medical University, with the approval of the Institutional Review Board for Clinical Research of The First Affiliated Hospital of Anhui Medical University. For the human blood samples used, the written informed consent was given to donors indicating that the blood left after the health examination was going to be used for research. Granulocytes were isolated using density gradient centrifugation with Percoll. To prepare the gradient, 2.5 ml of 75% Percoll was added to the bottom of a 15-ml centrifuge tube, followed by 2.5 ml of 62% Percoll layered on top. Fresh blood (1–3 ml) was diluted with serum-free RPMI 1640 to a total volume of 5 ml and carefully overlaid onto the 62% Percoll layer. The gradient was centrifuged at 200g for 25 min, followed by a second centrifugation at 400g for 15 min. Neutrophils were collected from the interface between the 62% and 75% Percoll layers.
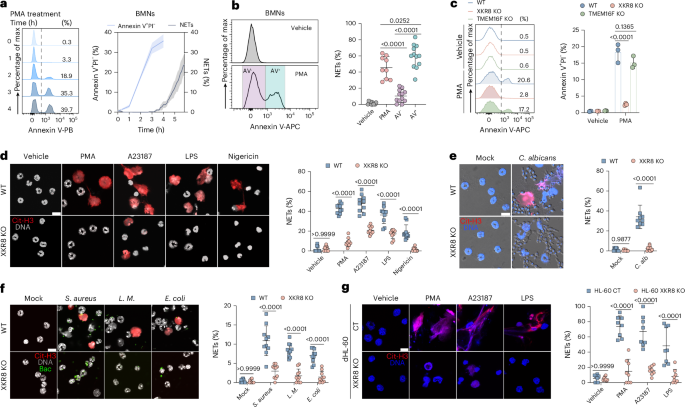
To assess NET formation and PS exposure, isolated neutrophils were treated with 100 nM PMA. Neutrophils were seeded either onto glass slides for immunofluorescence staining or into 96-well plates for flow cytometry. At the designated time points, cells on glass slides were fixed with 4% paraformaldehyde (PFA) for immunofluorescence staining. Alternatively, cells were collected, washed with PBS and stained with annexin V-APC along with either propidium iodide (1 μg ml−1) or DAPI (40 nM) for flow cytometry analysis.
NET induction and quantification
Freshly isolated neutrophils were stimulated immediately after purification with chemical reagents or microorganisms in serum-free RPMI 1640 medium supplemented with 10 mM HEPES. For chemical induction of NETs, neutrophils were treated with PMA (100 nM) for 6 h, LPS (10 μg ml−1) for 2 h or A23187 (1.25 μM), nigericin (5 μM), zymosan (25 μg ml−1) and MSU (100 μg ml−1) for 4 h in a cell culture incubator. For microbial induction, neutrophils (1.5 × 105 cells per well) were co-cultured with E. coli, L. monocytogenes, S. aureus or C. albicans at a MOI of 10 for 1.5 h. After stimulation, cells were fixed in 4% PFA for 10 min and permeabilized with 0.1% Triton X-100 in PBS for 20 min at room temperature. Fixed cells were blocked with 1% BSA and 10% goat serum or donkey serum for 1 h at room temperature before immunofluorescence staining. Neutrophils undergoing NET formation were labeled using antibodies against citrullinated histone H3 (Cit-H3, 1:1,000, ab5103, Abcam), MPO (1:1,000, AF3667, R&D system) or elastase (1:100, ab314916, Abcam), diluted in 1% BSA in PBS. NETs were visualized by mounting cells with DAPI and imaging using confocal microscopy (Olympus FV3000 and Nikon Ti2-E).
NETs were defined based on the DNA area observed in the fluorescence images. Semi-automated image analysis of DAPI-stained neutrophils was performed using Fiji software (v.2.14/1.54f), and calculated using MATLAB-based customer-written scripts ‘gating’ (Supplementary Materials 1), as outlined in Extended Data Fig. 1a. Image stacks were processed and converted in eight-bit grayscale. To quantify neutrophil numbers and measure the DNA area and fluorescence intensity of each cell, DAPI images were thresholded using ‘RenyiEntropy’ algorithm, followed by conversion to a mask and watershed segmentation to separate touching particles. NETs were recognized as structures with a DNA area greater than 80 μm2 in BMNs or 100 μm2 in dHL-60 cells. The percentage of NET formation was calculated by dividing the number of NETs by the total number of neutrophils in the field of view.
ROS measurements
ROS production was assessed using a luminol- and isoluminol-dependent chemiluminescence assay to measure intracellular and extracellular ROS, respectively, as previously described60. For intracellular ROS measurement following stimulation with PMA (100 nM), luminol (50 μM, A8515, Sigma-Aldrich) was added to the cell suspension along with membrane-impermeable superoxide dismutase (SOD; 50 U, S8160, Sigma-Aldrich) and catalase (2,000 U, C9322, Sigma-Aldrich) to eliminate extracellularly released O2− and H2O2. Chemiluminescence was recorded at 1-min intervals for 40 min at 37 °C using a BioTek Synergy HT microplate reader.
Extracellular ROS production was assessed by stimulating neutrophils with PMA in the presence of isoluminol (50 μM, A8264, Sigma-Aldrich) and exogenous horseradish peroxidase (HRP; 8 U, P8375, Sigma-Aldrich). The chemiluminescent signal was measured under the same conditions as described for intracellular ROS.
In addition, intracellular ROS levels were quantified using the fluorogenic probe dihydrorhodamine 123 (DHR123; 10 μM, C5313, APExBIO). Neutrophils were incubated with DHR123 for 30 min at 37 °C, followed by stimulation with PMA. The resulting fluorescence intensity was measured using flow cytometry, providing a quantitative assessment of intracellular ROS production.
To investigate the effects of ROS inhibitors or scavengers on NET formation, neutrophils were pretreated with NA-11 (1 μM), apocynin (30 μM), DPI (10 μM) or a dimethylsulfoxide (DMSO) vehicle control for 30 min before PMA (100 nM) stimulation for 6 h.
Evaluation of caspase-3 activation
Caspase-3 activation during the early stages of PMA-induced NET formation was evaluated using the CellEvent Caspase-3/7 Green Detection Reagent (C10423, Invitrogen) and an active caspase-3-specific antibody (570524, BD Biosciences). For the CellEvent assay, neutrophils were stained with CellEvent (1 μM) and subsequently stimulated with PMA (100 nM) in RPMI 1640 medium. Fluorescent signals were measured at defined time points by flow cytometry (LSR Fortessa, BD). Active caspase-3 was further detected using an antibody that specifically recognizes the cleaved form of caspase-3. Neutrophils undergoing NET formation were stained with Ly6G-AF488 (BioLegend, 127626) and Fixable Viability Dye eFluor 780 (eBioscience, 65-0865-18) for 15 min at room temperature to distinguish viable cells. Cells were then fixed with BD Cytofix Fixation Buffer (554655) and permeabilized with BD Perm/Wash buffer (554723). Subsequently, cells were stained with a rabbit anti-active caspase-3 antibody (1:1,000 dilution), followed by secondary staining with either goat anti-rabbit PE or F(ab’)2-goat anti-rabbit AF594 (1:1,000 dilution, A13203, Invitrogen). Stained samples were analyzed using a LSR Fortessa flow cytometer (BD) or sorted on an Aria III cell sorter (BD). Only eFluor 780-negative cells were sorted onto adhesive slides. These slides were then examined using a confocal microscope (Nikon Ti2-E) to visualize localization of activated caspase-3 during NET formation.
To investigate the role of caspase-3 on NET formation, mouse neutrophils were pretreated with caspase-3 inhibitors, including Q-VD (10 μM), Z-VAD (30 μM), emricasan (10 μM) or a DMSO vehicle control for 30 min, followed by PMA stimulation (100 nM) for 6 h at 37 °C. Human primary neutrophils were treated in the same manner, except that PMA stimulation was performed at 5 nM for 4 h. The impact of these inhibitors on NET formation was then assessed using confocal microscopy and flow cytometry.
Measurements of lipid scrambling
PS exposure was measured using annexin V staining. Following NET or apoptosis induction, neutrophils were pelleted and resuspended in 1× annexin V binding buffer (eBioscience), containing annexin V (5 ng ml−1) and either propidium iodide (100 ng ml−1, P4170, Sigma) or DAPI (40 nM, 422801, BioLegend). The samples were then analyzed using either a LSR Fortessa or a Symphony A5 flow cytometer (BD). To monitor lipid scrambling from the outer to the inner leaflet of PM, cells were stained with NBD-labeled phospholipid analogs, including NBD-phosphatidylcholine (NBD-PC; 810132P, Sigma) and NBD-sphingomyelin (NBD-SM; 810218 P, Sigma), as described previously38. After staining, an equal volume of cell suspension and prechilled RPMI1640 containing 5 mg ml−1 BSA was mixed to extract any unincorporated fluorescent lipid analogs. DAPI was added to exclude dead cells from the analysis. Additionally, PE exposure during lipid scrambling was evaluated using the PE-binding probe duramycin (Molecular Targeting Technologies). The stained samples were analyzed by flow cytometry on a LSR Fortessa flow cytometer (BD). Gating strategies are shown in Supplementary Fig. 3. Data from all experiments were processed and analyzed using FlowJo software v.10.
Live-cell imaging assay
To monitor chromatin area changes during NET formation, mouse bone-marrow-derived neutrophils were plated on a 35-mm glass-bottom imaging dish. After 30 min of incubation, attached cells were treated with 200 μl of RPMI 1640 with HEPES containing 20 μg ml−1 Hoechst 33342 (2031173, Invitrogen), 100 nM SYTOX Green (S7020, Invitrogen) and 100 nM PMA. For image analysis, regions of interest (ROIs) corresponding to imaged cells and Hoechst 33342-stained nuclei in each frame were automatically segmented by Cellpose algorithm using pretrained and custom-trained model ‘nucleus’. The nuclear area of each cell was extracted from the segmented ROIs and calculated using MATLAB-based custom-written scripts ‘NETs’ (Supplementary Materials 1). Time-lapse imaging data were analyzed using ImageJ, Cellpose v.2.2.3 and MATLAB (2023a).
Fluorescence and lifetime imaging
Fluorescence and lifetime imaging were performed on a laser confocal microscope (Leica TCS SP8X). For fluorescence lifetime imaging of Flipper-TR (CY-SC020, Cytoskeleton) in neutrophils, cells were first stained with Hoechst 33342 (20 μg ml−1), seeded in a four-chamber glass-bottom dish and treated with PMA (100 nM) at 37 °C in a 5% CO2 incubator for 2 h. Neutrophils were subsequently stained with Flipper-TR (1 μM), DAPI (40 nM) and Alexa Fluor 594-conjugated annexin V (A13203, Invitrogen) at 37 °C for 15 min before imaging. Lifetime imaging was conducted using Leica’s 480 nm pulsed laser for excitation, with photon collection in the 582–670 nm range. Only DAPI-negative cells were included in the subsequent analysis. For lifetime imaging analysis, the fluorescence lifetime (τ) of the cell membrane was measured, with analysis focused on the PM. Cells were categorized as annexin V+ or annexin V−, and the mean fluorescence lifetime was calculated using Leica software (Leica Application Suite X).
Calcium influx and cytosolic Ca2+ level detection
Calcium influx was detected using Indo-1 (65-0856-39, eBioscience). Calcium influx by Indo-1 was reported previously59. In brief, neutrophils were collected and resuspended in calcium-free HBSS containing 1 μM Indo-1, then incubated at 37 °C for 30 min. The cells were subsequently washed twice with calcium-free HBSS and resuspended in HBSS supplemented with 1 mM CaCl2 and MgCl2. Neutrophils were stimulated with LPS (10 μg ml−1), PMA (100 nM), A23187 (1.25 μM) or crosslinking CD16 (2.4G2, 553142, BD) with a secondary goat anti-rat IgG (112-005-062, Jackson ImmunoResearch). For calcium channel agonists experiments, WT and XKR8 KO neutrophils were loaded with Indo-1 and treated with probenecid (1 mM), GSK1016790A (100 nM), GSK1702934A (100 μM), Yoda1 (10 μM) or CBD (5 μM). Calcium influx was measured using flow cytometry for 5 min on a BD FACSymphony A5 flow cytometer, and calcium flux was quantified as the ratio of Indo-1 emission at 400 nm (bound Ca2+) to 475 nm (free Ca2+).
To evaluate intracellular Ca2+ levels during NET formation, 2 × 105 neutrophils labeled with Indo-1 were seeded in 96-well plates and induced by PMA (100 nM) in the presence or absence of agonists for 2 h at 37 °C in RPMI 1640. Subsequently cells were resuspended in 1× binding buffer containing annexin V (5 ng ml−1) and propidium iodide (100 ng ml−1) and analyzed on a BD Symphony A5. Neutrophils were gated into annexin V+ or annexin V− populations, and intracellular Ca2+ levels was determined from the ratio of bound to free Ca2+ Indo-1.
Phagocytosis and degranulation assay
Neutrophils were seeded into 96-well plates, and GFP-expressing E. coli or FITC-labeled latex beads (L4530, Sigma-Aldrich) were opsonized through preincubation in RPMI 1640 medium containing 10% mouse serum at 37 °C for 30 min. For the phagocytosis assay, 1 × 105 neutrophils per well were co-cultured with opsonized E. coli or beads at an MOI of 10 for 30 min in a cell culture incubator. After incubation, neutrophils were washed and stained with antibodies targeting CD11b (1:500 dilution, 101224, BioLegend) and Ly6G (1:500 dilution, 127608, BioLegend). Phagocytosis was quantified by flow cytometry (BD LSR Fortessa). For the degranulation assay, neutrophils were stained with anti-CD63 (1:500 dilution, 143906, BioLegend) following co-culture with bacteria. The mean fluorescence intensity of CD63 was measured as an indicator of neutrophil degranulation.
Mouse serum cytokine profiling
Whole blood samples from naive mice were collected and centrifuged at 3,000g for 20 min to obtain serum. Cytokine levels were quantified using a Luminex multiplex immunoassay (LX-MultiDTM-23, Bio-Rad).
Acute lung injury mouse models
ALI was induced in 6–8-week-old mice via intranasal administration of LPS (100 ng in 50 μl PBS per mouse). Mice receiving an equal volume of PBS served as unchallenged control group. At 24 h after treatment, BALF and lung tissue samples were collected. The acellular fraction of BALF was obtained through double centrifugation, and the supernatant was used for further analysis. dsDNA levels in the BALF were quantified using the dsDNA HS Assay kit (Q32851, Invitrogen).
For the C. albicans-induced ALI model, anesthetized mice were challenged with 1 × 106 or 3 × 106 (for survival monitoring) C. albicans in 50 μl PBS via the intranasal route. BALF was collected 24 h after infection and lung tissues were isolated and homogenized. Serial dilutions of the homogenates were plated on YPD agar plates and incubated at 37 °C for 24 h to quantify fungal burden. To evaluate the effect of GSK1016790A, Xkr8fl/flLy6gcre mice were anesthetized and then treated intranasally with 50 μl of 10 μM GSK1016790A before C. albicans infection.
For C. albicans systemic infection, mice were intravenously injected via the tail vein with 1 × 106 C. albicans (or 1.1 × 106 for survival studies) in 200 μl PBS. At 24 h after infection, the lungs, brain, liver and spleen were collected and homogenized. Homogenates were plated on YPD agar for 24 h to determine fungal burden. Portions of each organ were fixed in PFA for subsequent histopathological analysis.
To detect NETs in lung tissues, frozen and fixed lung sections (5-μm thick) were prepared for immunofluorescence staining. Sections were permeabilized in 0.5% Triton X-100 for 20 min and blocked in PBS containing 2% BSA and 3% of donkey serum (D9663, Sigma-Aldrich) for 1 h at RT. Samples were incubated overnight at 4 °C with anti-citrullinated histone H3 (1:1,000) and anti-MPO (1:1,000), followed by incubation with secondary antibodies (1 h at room temperature in the dark): goat anti-rabbit IgG-A488 (1:1,000, A11008, Invitrogen) and donkey anti-goat IgG-A647 (1:1,000, A32849, Invitrogen). Nuclei were counterstained with DAPI. Images were acquired using a Nikon confocal microscope equipped with a ×60 oil objective lens. For quantification of NETs, at least eight random fields per sample were analyzed. Image stacks were processed and converted to eight-bit grayscale. Each channel of images was thresholded using the ‘Default’ method in ImageJ, followed by conversion to masks, and the total area of Cit-H3-positive signal was measured. NETs formation was expressed as the ratio of Cit-H3 area to DAPI area in the field of view.
Collagen antibody-induced arthritis mouse model
To assess the role of XKR8 in autoimmune disease, a CAIA model was established using 8–9-week-old WT and XKR8 KO mice maintained under specific pathogen-free conditions. On day 0, mice received an intraperitoneal (i.p.) injection of 2.5 mg per mouse of anti-type II collagen 5-clone monoclonal antibody cocktail (Chondrex). On day 3 after antibody injection, to boost the inflammatory response, mice were administered an i.p. injection of LPS (50 μg in 100 μl PBS per mouse; Chondrex). Arthritis severity was monitored daily from day 3 to day 14 using a semiquantitative clinical scoring system (0–4 points per paw; maximum total score 16) based on the following criteria: 0, no visible swelling, erythema, or joint deformity; 1, localized swelling or erythema confined to the knuckle joints; 2, mild swelling extending to the ankle; 3, severe swelling involving the entire paw (digits to ankle); and 4, paw rigidity, joint deformation or impaired weightbearing. Hind paw thickness was measured daily with a digital vernier caliper.
Histological analyses of tissue damage
For histological analyses, the left lung lobes were removed and fixed in 4% neutral formaldehyde and embedded in paraffin for tissue sectioning. The embedded waxes were cut into 10-μm slices and flattened on glass slides, followed by rehydration and H&E staining. Evaluation of lung damage for LPS ALI model was performed using a reliable scoring system as previously reported44. To assess the severity of lung damage, five randomly chosen fields were evaluated according to five independent variables: neutrophils in the alveolar space, neutrophils in the interstitial space, hyaline membranes, proteinaceous debris filling airspaces and alveolar septal thickening. Each item was scored 0, 1 or 2 according to the severity, with differential weighting according to the relevance of these variables. The sum of the weighted variables was averaged according to the number of determined fields, yielding a final score between 0 and 10. Scoring of tissue damage for C. albicans infection was performed as previously described48, ten random images of each section were analyzed by fibrin deposition and bleeding in the tissue and scored 0, 1, 2 or 3 according to the severity. The sum of the variables was averaged according to the number of determined fields.
Immunoblot
Cells were lysed in RIPA buffer containing protease and phosphatase inhibitor cocktails (R0278, Sigma-Aldrich). Protein concentrations were determined using a BCA assay (71285-M, Sigma-Aldrich). Lysates were denatured at room temperature or by boiling for 10 min in 1× sample buffer. To assess phosphorylation of ERK and PKC, caspases or XKR8 (XKR8-GFP) activation, 30 μg of total protein per sample was resolved by SDS–PAGE and transferred onto PVDF membranes. Membranes were incubated overnight at 4 °C with the following primary antibodies: phospho-ERKT202/Y204 (1:1,000), phospho-PKC (1:1,000), caspase-3 (1:1,000), caspase-7 (1:1,000), caspase-8 (1:1,000), caspase-9 (1:1,000) and GFP-tag (1:2,000). After primary antibodies incubation, membranes were probed with HRP-conjugated secondary antibodies for 1 h at room temperature. Protein bands were visualized using an ECL substrate (34580, Thermo Scientific) and detected using the ChemiDoc XRS+ Imaging System (Bio-Rad).
Statistical analysis
For comparisons involving more than two groups, one-way ANOVA or two-way ANOVA with Tukey’s multiple-comparisons test was used. All statistical analyses were performed using GraphPad Prism (v.10). Each experiment was independently repeated at least three times, with detailed statistical information provided in the corresponding figure legends. n denotes the number of animals used, technical replicates or quantified images, as specified in the legends. Data are expressed as mean ± s.e.m. P < 0.05 was considered statistically significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.













Leave a Reply