Demographic characteristics of study participants
Considering the prevalence and clinical significance of co-infections with ADV (26.6%) and IAV (14.5%) among MPP patients (Supplementary Fig. 1), this study further collected the case information and BALF samples of the relevant patients for further analysis. A total of 73 MPP patients (35 males), aged between 2.25 and 12.25 years (mean age: 7.01 ± 2.44), were included in the study. Among them, 31 were categorized as belonging to the single MP group, while the remaining 42 cases were classified within the co-infection group. The co-infection group was further stratified into two subgroups: the co-ADV subgroup (n = 21), and the co-IAV subgroup (n = 21). Ten bronchial foreign bodies (BFB) patients without pulmonary inflammatory responses were enrolled in the control group.
There were no significant differences in age or gender distribution between the control group and the other groups. Compared to patients in the single MP group, those in the co-infection group exhibited significant differences in several clinical parameters, including hospital stays (p < 0.001), duration of fever (p = 0.008), white blood cell count (p = 0.002), neutrophil count (p = 0.001), macrophage count (p = 0.044), and procalcitonin (PCT) concentration (p = 0.016). Additionally, patients with co-infection exhibited a significantly higher rate of tachypnea (64.3%, p = 0.001), pulmonary necrosis (16.7%, p = 0.017), and extrapulmonary complications (21.4%, p = 0.037). Other clinical characteristics were presented in Table 1.
To further investigate the impact of co-infection on the severity of clinical symptoms in MPP patients, we conducted an analysis of the imaging data from these individuals. We measured and calculated the extent of pulmonary lesions, including consolidation and pleural effusion, in these patients. Our findings indicated that, while there was no significant difference in the degree of pleural effusion between the co-infection group and the single MP group, patients in the co-infection group demonstrated a markedly higher level of lung consolidation compared to those in the single MP group (p = 0.016) (Fig. 1A–C). According to the findings presented above, co-infections are associated with more severe clinical manifestations, poorer laboratory indicators, and worse disease outcomes in cases of MPP when compared to single MP infections.
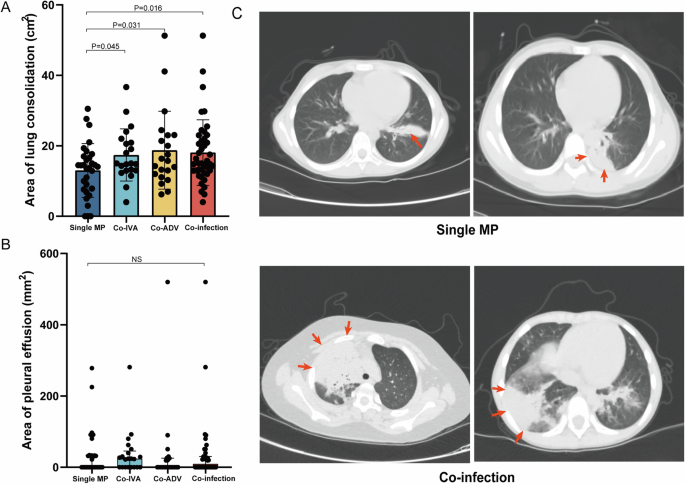
A Distribution of the size of lung consolidation area in patients among different groups; B Distribution of the size of pleural effusion area in patients between different groups; C Representative imaging findings from patients in different groups. Red arrows indicates the location and extent of lung consolidation. NS no significance, MP Mycoplasma pneumoniae, MPP Mycoplasma pneumoniae pneumonia, ADV adenovirus, IAV influenza A virus.
Metabolomic analysis identified pro-inflammatory metabolic patterns in BALF of MPP patients with co-infection
Principal component analysis (PCA) and cluster analysis identified significant differences in metabolite profiles within BALF samples between the MPP group and the control group (Supplementary Fig. 2A, B). A total of 296 differential metabolites (DMEs) were identified between the two groups (Supplementary Fig. 2C). The statistical significance of the observed fold changes in these DMEs is illustrated in a volcano plot, while the metabolic pathways enriched by these DMEs are elucidated through Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis (Supplementary Fig. 2D, E).
The results of the partial least squares discriminant analysis (PLS-DA) model demonstrated that, compared to the single MP group, the metabolic profiles of both the co-ADV and co-IAV groups exhibited significant differences. However, the metabolic patterns between the co-ADV and co-IAV groups were largely indistinguishable. Consequently, in subsequent metabolomic analyses, we directly compared the single MP group with the co-infection group (Fig. 2A, B). The clustering heatmap further validated this result (Fig. 2C). The results of the orthogonal projections to latent structures-discriminant analysis (OPLS-DA) model showed that R²Y = 0.972 and Q² = 0.855, indicating that the model had no risk of overfitting. In these two groups, a total of 2435 metabolites were identified, with 616 metabolites exhibiting significant differences in expression levels between the groups (Fig. 2D and Supplementary Data). These DMEs were displayed through a volcano plot (Fig. 2E).
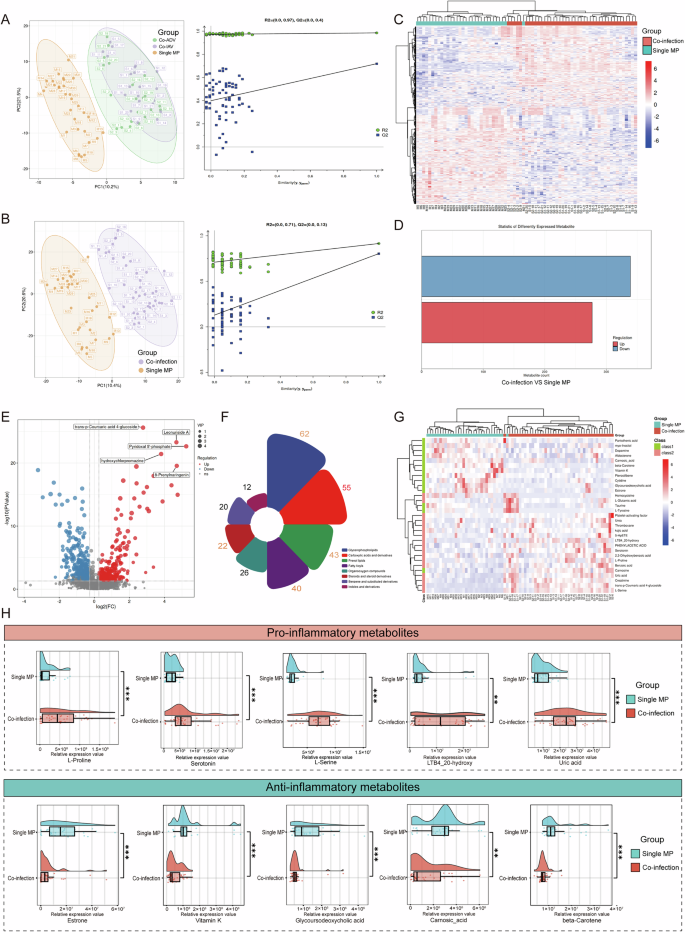
A, B Partial least squares discriminant analysis and permutation test among groups; C Clustering heatmap of pulmonary metabolites in MPP patients; D The number of differential metabolites; E The volcano plot clearly shows the types of metabolites that are significantly increased (red) or decreased (blue) in the co-infection group compared with the single MP group. Variable importance in projection (VIP) values were used to illustrate the contribution of metabolites to the orthogonal projections to latent structure-discriminant analysis model; F Classify the differential metabolites according to their types; G Cluster heatmap of differential metabolites with significant pro-inflammatory and anti-inflammatory biological functions. Class1 represents anti-inflammatory metabolites, and Class2 represents pro-inflammatory metabolites; H The distribution of differential metabolites with significant anti-inflammatory or pro-inflammatory effects between the two groups; **p < 0.01, ***p < 0.001.
Further classification of these DMEs revealed that the majority were identified as lipids, including glycerophospholipids, prenol lipids, fatty acyls, steroids, and steroid derivatives. Additionally, carboxylic acids and their derivatives were also prominently represented (Fig. 2F). We focused our attention on DMEs that exhibit well-defined pro-inflammatory and anti-inflammatory biological functions. Several well-recognized potent pro-inflammatory and anti-inflammatory mediators, along with their precursor molecules, exhibited significant differences between the groups (Fig. 2G). By integrating biological functions with fold change values, we identified the top 5 key pro-inflammatory and anti-inflammatory DMEs, as follows: the pro-inflammatory DMEs include L-Proline (p < 0.001, FC = 3.25), Serotonin (p < 0.001, FC = 2.94), L-Serine (p < 0.001, FC = 2.78), LTB4_20-hydroxy (p = 0.001, FC = 2.73), and Uric acid (p < 0.001, FC = 2.59); the anti-inflammatory DMEs include Estrone (p < 0.001, FC = 0.33), Vitamin K (p < 0.001, FC = 0.39), Glycoursodeoxycholic acid (p < 0.001, FC = 0.50), Carnosic acid (p = 0.001, FC = 0.55), and beta-Carotene (p < 0.001, FC = 0.57). The cloud and rain map clearly illustrated the expression patterns of key DMEs between the two groups. Specifically, in the co-infection group, the levels of pro-inflammatory metabolites were significantly elevated, while the levels of anti-inflammatory metabolites were markedly reduced. This observation elucidates the underlying reason for the more severe clinical manifestations observed in patients with co-infection (Fig. 2H).
Co-infection with ADV can specifically disrupt the lung microbiota structure in MPP patients
We initially analyzed the lung microbiota of the control group and all MPP patients. The results revealed significant differences in both Alpha (α)-diversity and Beta (β)-diversity between the two groups, suggesting a significant difference in the composition of the lung microbiota in MPP patients (Fig. 3A, B). The rarefaction curves indicate that the sequencing depth of the samples satisfies the criteria necessary for microbiota analysis (Supplementary Fig. 2F). We further identified the top 20 microbial species with the highest abundance in both groups at both the phylum and genus levels and conducted linear discriminant analysis (LDA) effect size (LEfSe) analysis to determine the differentially abundant species between the two groups (Fig. 3C, D).
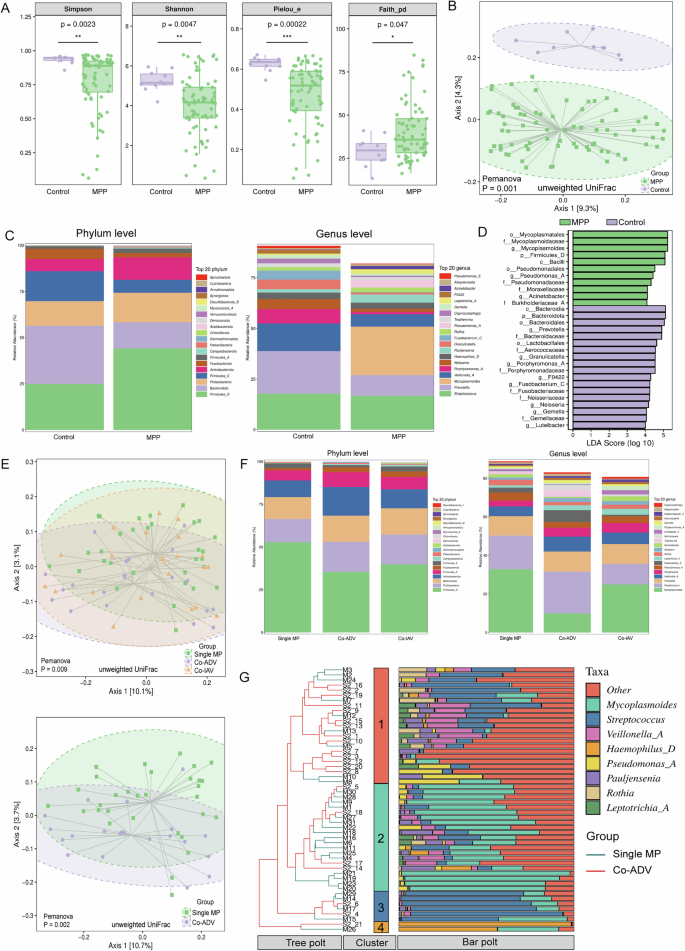
A, B Analysis of the α-diversity and β-diversity of the pulmonary microbiota in FBO patients and those with MPP. C Relative abundance of pulmonary microbiota at the phylum level and genus level in the two groups. D The LDA effect size analysis score identified differential microorganisms between the two groups. E Analysis of the β-diversity of the pulmonary microbiota in the Single MP group and co-infection groups. F Relative abundance of pulmonary microbiota at the phylum level and genus level in the three groups. G The results of cluster analysis of samples between the two groups. *p < 0.05, **p < 0.01, ***p < 0.001.
We further focused on the differences between the single MP group and the co-infection group. The findings indicated that although no significant difference was observed in the α-diversity index across these groups, a significant difference was detected in the β-diversity. This distinction was particularly pronounced between the single MP group and the co-ADV group (Fig. 3E). The lung exhibited comparable microbial compositions among three groups at the phylum level, but at the genus level, there were significant differences in the composition of lung microbiota among the three groups of patients (Fig. 3F). Specifically, the relative abundance of Streptococcus in the microbiota of BALF samples from patients in the co-ADV group was markedly higher compared to the other two groups. Compared to single MP infection, which accounts for 32.8%, co-infection—particularly with ADV, observed at a rate of 9.91%—led to a reduction in the relative abundance of MP in the BALF of patients with MPP. This phenomenon may be attributed to competitive interactions between the pathogens involved. Therefore, in the subsequent microbiota analysis, we further focused on the co-ADV group and the single MP group. All samples within these two groups were subjected to cluster analysis and divided into four clusters. The results revealed that a higher proportion of co-infection patients were classified into Cluster 1, which was predominantly characterized by a relatively high prevalence of other bacterial species (14/23). In contrast, patients with single MP infection were primarily categorized into Cluster 2, which was dominated by MP (17/21), further substantiating the aforementioned conclusion (Fig. 3G). Additionally, Cluster 3 and Cluster 4 were characterized by microbial profiles predominantly associated with Streptococcus and Haemophilus_D, respectively. However, the distribution of patients across different groups within these two clusters did not exhibit any statistically significant differences.
We subsequently investigated which microorganisms’ differential distributions predominantly contributed to these variations. Firstly, a Venn diagram was employed to elucidate the distinct distribution patterns of microbiota across diverse groups (Fig. 4A). Secondly, LEfSe and random forests (RF) models were used to analyze the differences in microbiota between single MP groups and co-ADV group, as well as to identify potential differential microorganisms (DMIs) (Fig. 4B–D). In comparison to the single MP group, the lung microbiota of patients in the co-ADV group experienced a reduction in beneficial bacteria (e.g., Lactobacillus) and an increase in anaerobic and opportunistic pathogenic microorganisms (such as Actinomyces and Leptotrichia_A). The distribution of these important DMIs in the samples from both groups was further clarified (Fig. 4E), and their classification ability was validated through receiver operating characteristic curves (ROCs) analysis (Fig. 4F). The findings indicated significant differences in the abundance of four microorganisms between the two groups. Notably, Lactobacillus [area under the curve (AUC) = 0.81, 95% confidence interval (CI): 0.69–0.94] may serve as a significant indicator bacterium for distinguishing between the single MP group and the co-ADV group.
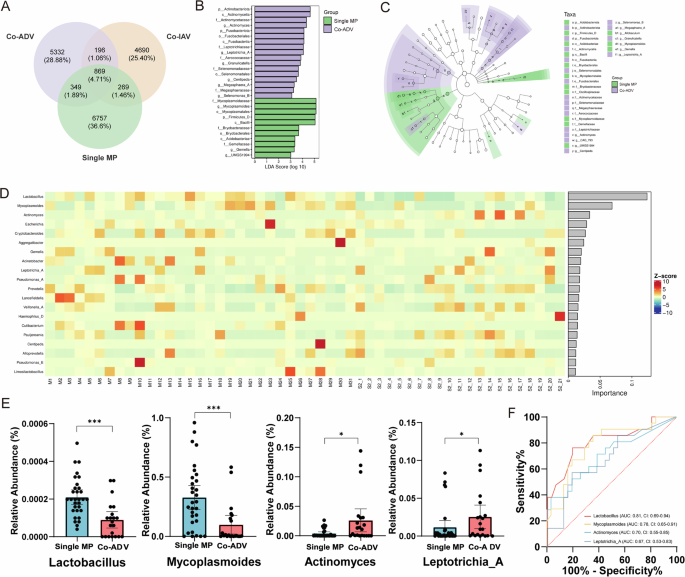
A Venn diagram showing the number of unique or shared microbial ASVs among different groups; B, C The LDA effect size analysis score and the effect size analysis of linear discriminant analysis identified differential microorganisms between the two groups; D Random Forest model identified the top 20 most influential marker microorganisms between the two groups, and provided importance scores for each microorganism in the model; E Distribution of differential microorganisms between the two groups; F ROC curves were constructed to study the diagnostic value of the differential microorganisms for ADV co-infection. AUC area under the curve, CI 95% confidence interval, MP Mycoplasma pneumoniae, ADV adenovirus, IAV influenza A virus. *p < 0.05, **p < 0.01, ***p < 0.001.
Co-infection with ADV can specifically enhance the expression of CCL family chemokines in MPP patients
Through the application of protein array technology, this study first analyzed the characteristics of the cytokine profiles in the BALF of BFB patients as well as all MPP patients. The results showed that the concentration of interleukin (IL)-17A in most patients was below the detection limit, and we were unable to conduct an effective analysis of this indicator. In addition, except for IL-5, IL-8, granulocyte colony-stimulating factor (G-CSF), and platelet-derived growth factor BB (PDGF-BB), the distributions of the remaining 20 cytokines in the two groups were significantly different (Supplementary Fig. 3A). This finding suggests that patients with MPP may undergo a significant “cytokine storm.” This phenomenon could be closely associated with the case selection criteria employed in this study, as all cases necessitated alveolar lavage.
Further research results demonstrated that, compared with the single MP group, all CCL family chemokines and PDGF-BB were significantly upregulated in the co-ADV group (Fig. 5A). Subsequently, we utilized the STRING database (https://cn.string-db.org/) to perform an interaction analysis of the differentially expressed cytokines and conducted a Gene Ontology Biological Process enrichment analysis (Fig. 5B, C). Our findings indicated that these differential proteins were significantly associated with monocyte chemotaxis. We further conducted a correlation analysis between the concentrations of differential cytokines (DCKs) and peripheral blood monocyte counts in patients. The results showed that the concentrations of monocyte chemotactic protein-1 (MCP-1) (r = 0.26, p = 0.024), macrophage inflammatory protein-1α (MIP-1α) (r = 0.27, p = 0.019), and MIP-1β (r = 0.24, p = 0.044) were positively correlated with the number of peripheral blood monocytes (Fig. 5D and Supplementary Fig. 3B). Therefore, co-infection with ADV can specifically enhance the expression of CCL family chemokines in the lungs of MPP patients, and this phenomenon is associated with an increase in peripheral blood monocyte counts. However, further research is required to confirm the causal relationship between these factors.
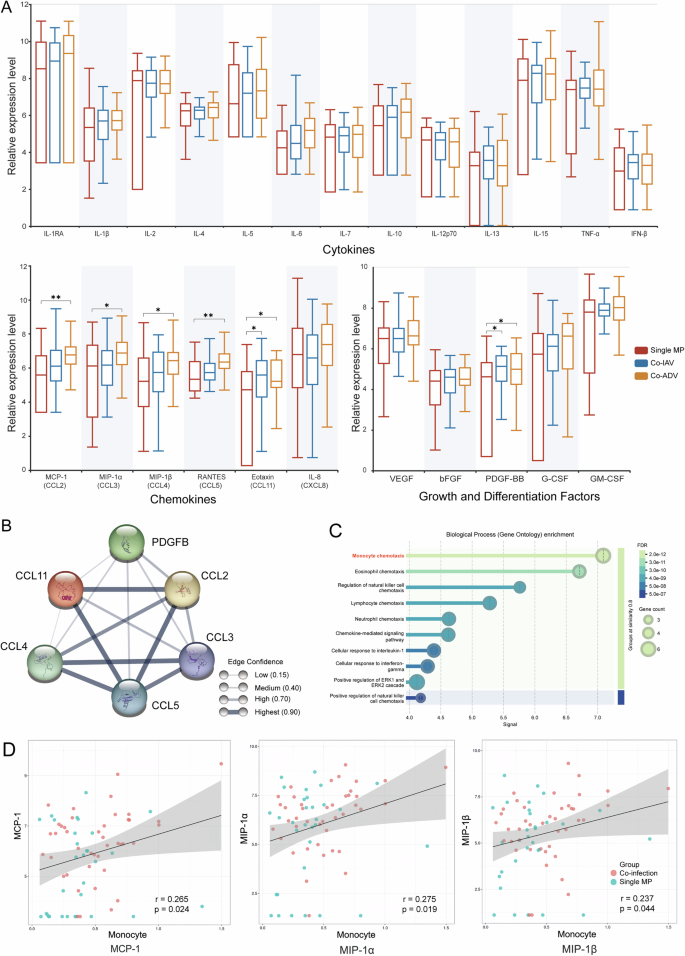
A The distribution of cytokines between the co-infection group and the single MP group; B Analysis of the interactions between differentially expressed cytokines; C Gene Ontology Biological Process enrichment analysis of differentially expressed cytokines; D Correlation analysis between differential cytokines and peripheral blood monocyte. *p < 0.05, **p < 0.01.
The intricate interactions and associations between the microbiome, metabolome, cytokine profile, and host immune response
To investigate the association between the lung microbiome and metabolome in MPP patients with, we constructed an association network that linked DMIs and DMEs in patients from both the single MP group and the co-infection group. In this network, on average, 9.8 DMEs are associated with each DMI (Fig. 6A). Among these, Lactobacillus and Limosilactobacillus exhibited significant correlations with 41 and 28 DMEs, respectively, sharing 14 identical metabolites. Additionally, the zi-pi value highlights key nodes within the network. The results indicate that Limosilactobacillus is identified as a network hub (Fig. 6B). These findings suggest that the depletion of beneficial microorganisms may be a critical factor in the pathogenesis of co-infected MPP, and their functions show a high degree of similarity.
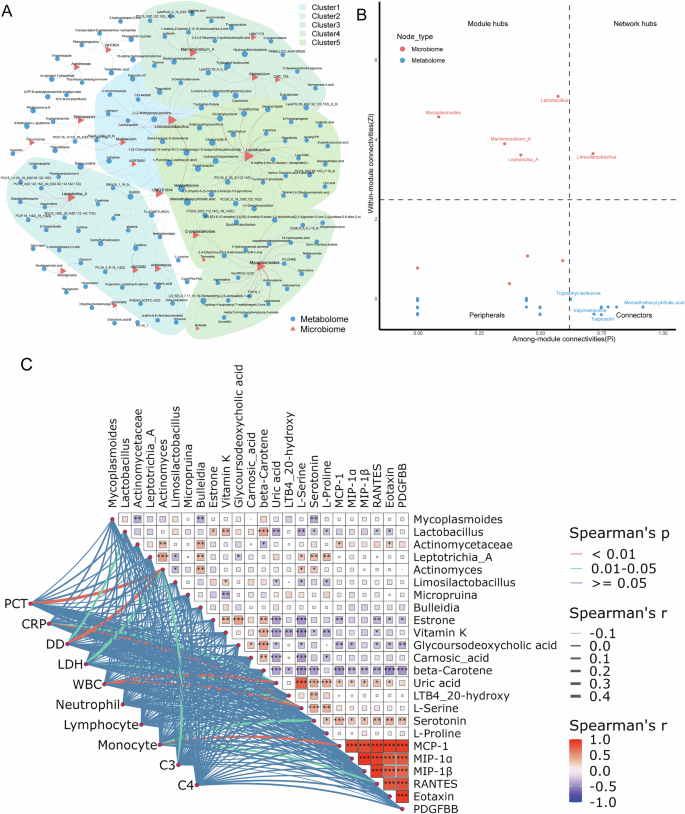
A Network clustering analysis of differential microorganisms and metabolites between the single MP group and the co-infected group; B The Zi-Pi map shows the topological properties of important nodes in the network clustering analysis. C Network heatmaps elucidate the correlations between key clinical indicators and major differential microorganisms, metabolites, and cytokines. *p < 0.05, **p < 0.01, ***p < 0.001.
We further revealed the potential interactions among different omics, cytokine profiles, and clinical indicators between the co-infection group and the single MP group through network heatmaps. The results demonstrated significant correlations among DMIs, DMEs, DCKs, and the clinical indicators of patients (Fig. 6C). Overall, our results suggest that these differential substances are significantly associated with the clinical indicators of patients and could potentially reflect the disease severity of MPP.














Leave a Reply