Noy, R. & Pollard, J. W. Tumor-associated macrophages: from mechanisms to therapy. Immunity 41, 49–61 (2014).
Pittet, M. J., Michielin, O. & Migliorini, D. Clinical relevance of tumour-associated macrophages. Nat. Rev. Clin. Oncol. 19, 402–421 (2022).
Allavena, P., Anfray, C., Ummarino, A. & Andón, F. T. Therapeutic manipulation of tumor-associated macrophages: facts and hopes from a clinical and translational perspective. Clin. Cancer Res. 27, 3291–3297 (2021).
Barry, S. T., Gabrilovich, D. I., Sansom, O. J., Campbell, A. D. & Morton, J. P. Therapeutic targeting of tumour myeloid cells. Nat. Rev. Cancer 23, 216–237 (2023).
Franklin, R. A. et al. The cellular and molecular origin of tumor-associated macrophages. Science 344, 921–925 (2014).
Locati, M., Curtale, G. & Mantovani, A. Diversity, mechanisms, and significance of macrophage plasticity. Annu. Rev. Pathol. 15, 123–147 (2020).
Schaer, D. J. et al. Hemorrhage-activated NRF2 in tumor-associated macrophages drives cancer growth, invasion, and immunotherapy resistance. J. Clin. Invest. 134, e174528 (2023).
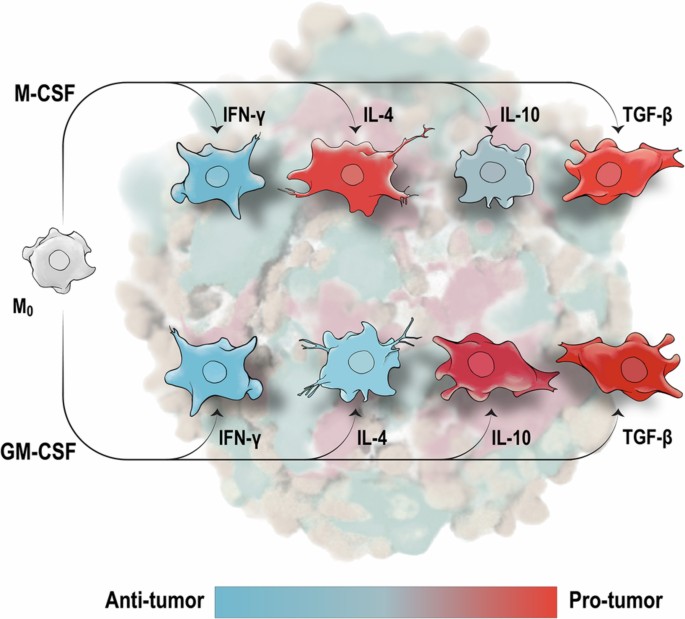
Mantovani, A., Sozzani, S., Locati, M., Allavena, P. & Sica, A. Macrophage polarization: tumor-associated macrophages as a paradigm for polarized M2 mononuclear phagocytes. Trends Immunol. 23, 549–555 (2002).
Murray, P. J. Macrophage polarization. Annu. Rev. Physiol. 79, 541–566 (2017).
Ginhoux, F., Schultze, J. L., Murray, P. J., Ochando, J. & Biswas, S. K. New insights into the multidimensional concept of macrophage ontogeny, activation and function. Nat. Immunol. 17, 34–40 (2016).
Sanin, D. E. et al. A common framework of monocyte-derived macrophage activation. Sci. Immunol. 7, eabl7482 (2022).
Becher, B., Tugues, S. & Greter, M. GM-CSF: from growth factor to central mediator of tissue inflammation. Immunity 45, 963–973 (2016).
Achuthan, A. et al. Granulocyte macrophage colony-stimulating factor induces CCL17 production via IRF4 to mediate inflammation. J. Clin. Invest. 126, 3453–3466 (2016).
Gordon, S. Alternative activation of macrophages. Nat. Rev. Immunol. 3, 23–35 (2003).
Hamilton, J. A. Colony-stimulating factors in inflammation and autoimmunity. Nat. Rev. Immunol. 8, 533–544 (2008).
Hamilton, J. A., Cook, A. D. & Tak, P. P. Anti-colony-stimulating factor therapies for inflammatory and autoimmune diseases. Nat. Rev. Drug Discov. 16, 53–70 (2016).
Helft, J. et al. GM-CSF mouse bone marrow cultures comprise a heterogeneous population of CD11c(+)MHCII(+) macrophages and dendritic cells. Immunity 42, 1197–1211 (2015).
Vallelian, F. et al. Heme-stress activated NRF2 skews fate trajectories of bone marrow cells from dendritic cells towards red pulp-like macrophages in hemolytic anemia. Cell Death Differ. 29, 1450–1465 (2022).
Bill, R. et al. CXCL9:SPP1 macrophage polarity identifies a network of cellular programs that control human cancers. Science 381, 515–524 (2023).
Rodriguez, P. C. et al. Arginase I production in the tumor microenvironment by mature myeloid cells inhibits T-cell receptor expression and antigen-specific T-cell responses. Cancer Res. 64, 5839–5849 (2004).
Matsubara, E. et al. SPP1 derived from macrophages is associated with a worse clinical course and chemo-resistance in lung adenocarcinoma. Cancers 14, 4374 (2022).
Schaer, D. J. et al. Stress-NRF2 response axis polarizes tumor macrophages and undermines immunotherapy. J. Immunother. Cancer 13, e013063 (2025).
LaMarche, N. M. et al. An IL-4 signalling axis in bone marrow drives pro-tumorigenic myelopoiesis. Nature 625, 166–174 (2024).
Mollaoglu, G. et al. Ovarian cancer-derived IL-4 promotes immunotherapy resistance. Cell 187, 7492–7510.e22 (2024).
Tel, J. et al. Tumoricidal activity of human dendritic cells. Trends Immunol. 35, 38–46 (2014).
Ferris, S. T. et al. cDC1 prime and are licensed by CD4+ T cells to induce anti-tumour immunity. Nature 584, 624–629 (2020).
Kang, S. et al. Engineered GM-CSF polarizes protumorigenic tumor-associated macrophages to an antitumorigenic phenotype and potently synergizes with IL-12 immunotherapy. J. Immunother. Cancer 12, e009541 (2024).
Ruffolo, L. I. et al. GM-CSF drives myelopoiesis, recruitment and polarisation of tumour-associated macrophages in cholangiocarcinoma and systemic blockade facilitates antitumour immunity. Gut 71, 1386–1398 (2022).
Duluc, D. et al. Interferon-gamma reverses the immunosuppressive and protumoral properties and prevents the generation of human tumor-associated macrophages. Int. J. Cancer 125, 367–373 (2009).
Dangaj, D. et al. Cooperation between constitutive and inducible chemokines enables T cell engraftment and immune attack in solid tumors. Cancer Cell 35, 885–900.e10 (2019).
Bronte, V. & Murray, P. J. Understanding local macrophage phenotypes in disease: modulating macrophage function to treat cancer. Nat. Med. 21, 117–119 (2015).
Kumar, A., Taghi Khani, A., Sanchez Ortiz, A. & Swaminathan, S. GM-CSF: A double-edged sword in cancer immunotherapy. Front. Immunol. 13, 901277 (2022).
Hong, I.-S. Stimulatory versus suppressive effects of GM-CSF on tumor progression in multiple cancer types. Exp. Mol. Med. 48, e242–e242 (2016).
Henriques, A. et al. TGF-β builds a dual immune barrier in colorectal cancer by impairing T cell recruitment and instructing immunosuppressive SPP1 macrophages. Nat. Genet. 57, 3050–3065 (2025).
Kumar, V. et al. Cancer-associated fibroblasts neutralize the anti-tumor effect of CSF1 receptor blockade by inducing PMN-MDSC infiltration of tumors. Cancer Cell 32, 654–668.e5 (2017).
Pfefferlé, M. et al. Antibody-induced erythrophagocyte reprogramming of Kupffer cells prevents anti-CD40 cancer immunotherapy-associated liver toxicity. J. Immunother. Cancer 11, e005718 (2023).
Anderson, N. M. & Simon, M. C. The tumor microenvironment. Curr. Biol. 30, R921–R925 (2020).
Humar, R., Schaer, D. J. & Vallelian, F. Erythrophagocytes in hemolytic anemia, wound healing, and cancer. Trends Mol. Med. 28, 906–915 (2022).
Pfefferlé, M. et al. Hemolysis transforms liver macrophages into antiinflammatory erythrophagocytes. J. Clin. Invest. 130, 5576–5590 (2020).
Lavin, Y. et al. Tissue-resident macrophage enhancer landscapes are shaped by the local microenvironment. Cell 159, 1312–1326 (2014).
Daniel, B. et al. Macrophage inflammatory and regenerative response periodicity is programmed by cell cycle and chromatin state. Mol. Cell 83, 121–138.e7 (2023).
Na, Y. R. et al. GM-CSF induces inflammatory macrophages by regulating glycolysis and lipid metabolism. J. Immunol. 197, 4101–4109 (2016).
Ryan, D. G. & O’Neill, L. A. J. Krebs cycle reborn in macrophage immunometabolism. Annu. Rev. Immunol. 38, 289–313 (2020).
Chen, C. et al. Epigenetic regulation of macrophage polarization in wound healing. Burns Trauma 11, tkac057 (2023).
Park, M. D., Silvin, A., Ginhoux, F. & Merad, M. Macrophages in health and disease. Cell 185, 4259–4279 (2022).
McNelis, J. C. & Olefsky, J. M. Macrophages, immunity, and metabolic disease. Immunity 41, 36–48 (2014).
Eming, S. A., Murray, P. J. & Pearce, E. J. Metabolic orchestration of the wound healing response. Cell Metab. 33, 1726–1743 (2021).
Liao, Y., Smyth, G. K. & Shi, W. The R package Rsubread is easier, faster, cheaper and better for alignment and quantification of RNA sequencing reads. Nucleic Acids Res. 47, e47 (2019).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Lun, A. T. L., McCarthy, D. J. & Marioni, J. C. A step-by-step workflow for low-level analysis of single-cell RNA-seq data with Bioconductor. F1000Res 5, 2122 (2016).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Badia-I-Mompel, P. et al. decoupleR: ensemble of computational methods to infer biological activities from omics data. Bioinform. Adv. 2, vbac016 (2022).
Vallelian F. An ontogeny-cytokine code determines macrophage response polarity and tumor outcomes – Sequencing data accession number GSE304868 (Gene Expression Omnibus; https://www.ncbi.nlm.nih.gov/geo/).













Leave a Reply