Human PBMC-derived macrophages obtention and cell culture
Peripheral blood mononuclear cells (PBMC) were separated from buffy coats (obtained from Etablissement Français du Sang) using a Ficoll gradient method. Briefly, blood was diluted half in PBS, added on Ficoll (CMSMSL01-01, Eurobio Scientific, Les Ulis, France) (2:1 v:v) and centrifugated 20 min at 700 g with low acceleration and no braking. Then, separated PBMC were washed and remaining red cells were lysed (lysis buffer: 150 mM NH4Cl, 10 mM KHCO3 and 0,1 mM EDTA). PBMC were seeded and incubated 1 h in culture conditions (i.e. 5% CO2, 37 °C). After adhesion, monocytes were differentiated in M2 macrophages with 100 ng/mL macrophage colony stimulating factor (M-CSF, 130-096-492, Miltenyi, Bergisch Gladbach, Germany) in RPMI medium with 10% heat inactivated FBS during 6 days, with differentiation medium renewal after 3 days. Macrophages were then activated with 20 ng/mL interleukine-4 (IL-4, 130-093-922, Miltenyi) in serum free Opti-MEM medium for 24 h. The GRP94 inhibitor PU-WS13 (HY-18680, MedChemExpress, Monmouth Junction, New Jersey, USA) was added 24 h before activation and during the whole activation period.
Human TNBC cells MDA-MB-231 (HTB-26™, ATCC, Manassas, VA, USA) were grown in DMEM with 10% heat inactivated FBS in previously mentioned culture conditions (i.e., 5% CO2, 37 °C) and tested for mycoplasma contamination before use. For experiments, cells were seeded 24 h in growth conditions before being treated with GRP94 inhibitors, PU-WS13 or GRP94 inhibitor-1 (HY-112910, MedChemExpress), during 24 h in serum free DMEM.
Binding affinity assay by microscale thermophoresis (MST)
Specific binding between recombinant human furin (1503-SE, R&D Systems®, USA) and recombinant human GRP94 (RD172379100, Biovendor®, Czech Republic) was measured by MST assays. Briefly, recombinant furin was serially diluted in PBS-Tween 20 0.01% buffer from a maximal furin concentration at 3.58 μM to 0.1 nM and Nano-RED-labelled recombinant GRP94 was added at a constant concentration of 100 nM for each furin dilution point. A negative control was also performed with the same procedure by replacing furin with a recombinant human Hsp110 kindly provided by Dr. François Hermetet (Inserm UMR 1231, Dijon, France) diluted from 2.33 μM to 0.07 nM. The mixtures were loaded into Monolith NT.115 Capillaries (NanoTemper Technologies, Germany) and thermophoresis was performed using a Monolith NT.115 (NanoTemper Technologies, Germany) at room temperature with excitation power and MST power both at 40%. The data were analyzed using MO. Affinity Analysis v2.3 software (NanoTemper Technologies, Germany).
Furin enzymatic activity assay
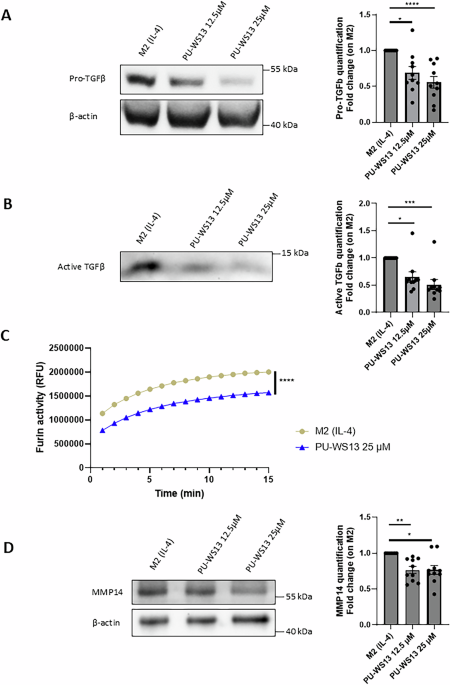
After treatment, PBMC-derived M2 macrophages or MDA-MB-231 cells were harvested by scrapping, washed with PBS and lysed using a TRITON X-100 based lysis buffer (20 mM Tris-HCl, 150 mM NaCl, 1% Triton X-100, H2O). Cell lysate (20 μL) was diluted in 20 μL of ultrapure H2O and 1 nmol of a fluorogenic substrate to furin: Pyr-Arg-Thr-Lys-Arg-AMC trifluoroacetate salt (4018149, Bachem, Bubendorf, Switzerland). Furin enzymatic activity was determined by recording fluorescence (355 nm excitation and 460 nm emission) over time using an EnVision® multimode plate reader (PerkinElmer®, Waltham, Massachusetts, USA).
Western blotting
PBMC-derived M2 macrophages or MDA-MB-231 cells were harvested by scrapping, washed with PBS and lysed 20 min at 4 °C in cell lysis buffer (#9803, Cell Signaling Technology, Danvers, Massachusetts, USA) supplemented with Complete™ Protease Inhibitor Cocktail (11697498001, Merck, Darmstadt, Germany) and phosphatase inhibitor cocktails 2 and 3 (P5726 and P0044, Sigma-Aldrich, Saint-Louis, Missouri, USA). After centrifugation 15 min at 14,000 g to remove cellular debris, samples were prepared in classical loading buffer, heated 5 min at 95 °C, loaded on SDS-PAGE gels before being transferred on PVDF membranes and saturated with 3% BSA diluted in TBS-0,1% Tween-20. PVDF membranes were incubated with primary antibodies (diluted at 1:1000 in saturation buffer) overnight under constant agitation at 4 °C and, after washing, with the corresponding HRP-conjugated secondary antibody (712-036-153, Jackson Immunol Research, Cambridge, UK; #7074, Cell Signaling Technology, Danvers, Massachusetts, USA; 1:10000 dilution in saturation buffer) 1 h at room temperature. Primary antibodies: GRP94 (ADI-SPA-850, Enzo Life Sciences, Villeurbanne, France), TGF-β1 (ab215715, Abcam, Cambridge, UK), Furin (ab183495, Abcam), MMP14 (AB6004, Merck). Mouse monoclonal Anti-β-Actin−Peroxidase antibody (A3854, Sigma-Aldrich) was used for loading control.
For western blotting on supernatants, cell culture supernatants were collected, centrifugated 5 min at 400 g to remove remaining cells and concentrated using Amicon® Ultra-0.5 Centrifugal Filter Devices with a 3 kDa molecular weight cut-off (UFC500396, Merck) following the manufacturer’s instructions. Briefly, 500 µL supernatants were added on filter devices, centrifugated 30 min at 14,000 g and finally turned upside down in clean tubes and centrifugated again 2 min at 1000 g to collect concentrated supernatants. Samples were then prepared for SDS-PAGE western blotting as described above.
GRP94-Furin co-immunoprecipitation assay
PBMC-derived M2 macrophages or MDA-MB-231 cells were harvested by scrapping, washed with PBS and lysed 20 min at 4 °C in cell lysis buffer (#9803, Cell Signaling Technology, Danvers, Massachusetts, USA) supplemented with Complete™ Protease Inhibitor Cocktail (11697498001, Merck, Darmstadt, Germany). After centrifugation 15 min at 14,000 g to remove cellular debris, samples were incubated overnight at 4 °C with 4 µg of anti-GRP94 (ab3674, Abcam) or a non-relevant antibody (X0903, Dako, Santa Clara, California, USA). Then, samples were incubated 30 min at 4 °C with 100 µL of µMACS™ Protein A MicroBeads (130-071-001, Miltenyi) and loaded on µColumns (130-042-701, Miltenyi) placed on a magnetic µMACS™ Separator (130-042-602, Miltenyi). Samples were washed 4 times with lysis buffer, incubated 5 min with 20 µL loading buffer heated at 95 °C and eluted using 50 µL of hot loading buffer. Finally, samples were heated 5 min at 95 °C and SDS-PAGE western blotting was performed as described above. Anti-vinculin antibody (V9131, Merck) was used for input loading control.
Immunofluorescence
PBMC-derived M2 macrophages seeded on coverslips were washed with PBS, fixed with a 4% formaldehyde solution for 15 min and washed again with PBS. Autofluorescence was then quenched with 50 mM NH4Cl for 10 min. After being washed, cells were permeabilized with PBS – 0.1% TRITON X-100, blocked for 1 h with PBS-1% BSA and incubated with primary antibodies diluted at 1:100 in PBS-1% BSA at 4 °C overnight: anti-GRP94 (ab210960, Abcam) and anti-LRRC33 (PA5-23822, Invitrogen, Waltham, Massachusetts, USA). Then cells were washed and stained 30 min at room temperature with secondary fluorescent antibodies (A11008 and A11004, Invitrogen) diluted at 1:1000 in PBS-1% BSA. After washes, coverslips were mounted in ProLong® Gold Antifade Reagent with DAPI (P36941, ThermoFisher Scientific, Waltham, Massachusetts, USA). Microscopy images were acquired using an Axio Imager 2 equipped with an AxioCam MRm monochrome CCD camera (Carl Zeiss Microscopy GmbH, Jena, Germany) and analyzed with Zen 2.3 Lite® software (Carl Zeiss Microscopy GmbH, Germany).
Duolink® Proximity-Ligation Assay (PLA™)
PLA™ was performed with Duolink® In Situ Detection Reagents Orange kit (DUO92007, Sigma-Aldrich). Cells seeded on coverslips were washed twice with PBS and fixed in 4% formaldehyde solution for 10 min at 4 °C. They were then permeabilized and saturated with a PBS – 3% BSA – 0.2% saponin solution for 20 min at room temperature before being incubated overnight at 4 °C with primary antibodies couples diluted at 1:100 in saturation buffer: anti-GRP94 (ab210960 or ab3674, Abcam) and one of the following antibodies: anti-Furin (ab183495, Abcam), anti-LRRC33 (PA5-23822, Invitrogen) or anti-HSP105 (sc-74550, SantaCruz Biotechnology, Dallas, Texas, USA). After incubation, cells were washed with PBS and incubated with Duolink® In Situ PLA® Probe Anti-Rabbit PLUS (DUO92002, Sigma-Aldrich) and Duolink® In Situ PLA® Probe Anti-Mouse MINUS (DUO92004, Sigma-Aldrich) for 1 h at 37 °C. Thereafter, PLA™ protocol was performed following the manufacturer’s protocol. Finally, slides were mounted in ProLong® Gold Antifade Reagent with DAPI (P36941, ThermoFisher Scientific). Microscopy images were acquired using an Axio Imager 2 equipped with an AxioCam MRm monochrome CCD camera (Carl Zeiss Microscopy GmbH) and analyzed by Zen 2.3 Lite® software (Carl Zeiss Microscopy GmbH). Spots quantification was performed using Icy® 2.0 software (BioImage Analysis Lab, Institut Pasteur, France).
MDA-MB-231 cell migration analysis using wound healing assay
MDA-MB-231 cells were seeded in a 96-wells plate and cultured during 24 h in growth conditions, then starved with DMEM – 0,5% FBS during 24 h. After starving, the wound was realized using an Incucyte® WoundMaker 96-Tool (4563, Sartorius, Göttingen, Germany). Then, cells were washed with HBSS and treatments with the GRP94 inhibitors, PU-WS13 or GRP94 inhibitor-1, added in serum free DMEM. Finally, cell migration in wound areas was measured during 24 h in culture conditions (5% CO2, 37 °C) using an Incucyte® S3 Live-Cell Analysis System (Sartorius) and relative wound densities over time calculated using Incucyte® Scratch Wound Analysis Software Module (9600-0012, Sartorius).
Cytotoxicity assay
Cytotoxicity of PU-WS13 and GRP94-inhibitor-1 on human primary M2 macrophages and MDA-MB-231 cells was assessed using the MTS Cell Proliferation Assay Kit (ab197010, Abcam) following the manufacturer’s instructions. Cells treated with 10 or 20% DMSO, respectively for MDA-MB-231 or M2 macrophages, were used as positive controls.
TGF-β1 and MMP14 mRNA transcription analysis by RT-qPCR
After treatment, PBMC-derived M2 macrophages total RNA was extracted using ReliaPrep RNA Miniprep Systems (Z6011, Promega, Madison, Wisconsin, USA). Reverse transcription was performed using the Maxima First Strand cDNA Synthesis Kit for RT-qPCR, with dsDNase (#K1672, ThermoFisher Scientific), and qPCR was performed using the iTaq Universal SYBR Green Supermix (1725121, Bio-Rad, Hercules, California, USA) in a ViiA 7 Real-Time PCR System (ThermoFisher). Experiments were performed following manufacturer’s instructions. Primers for qPCR were purchased from Qiagen (Hilden, Germany): Hs_GAPDH_1_SG QuantiTect Primer Assay (QT00079247, 249900), Hs_TGFB1_1_SG QuantiTect Primer Assay (QT00000728, 249900), and Hs_MMP14_1_SG QuantiTect Primer Assay (QT00001533, 249900).
Statistical analysis
Statistical analysis of data was performed using GraphPad Prism 10.2.3 software (La Jolla, CA, USA). Shapiro-Wilk test was used to assess data compliance with normal homogeneity of variance. Western-blot quantification and cytotoxicity data were analyzed through One-way ANOVA with Friedman test using the two-stage linear step-up procedure of Benjamini, Krieger and Yekutieli. PLA spots quantification data were analyzed using two-tailed Mann-Whitney test. RT-qPCR fold increase data were analyzed using two-tailed Wilcoxon test. Two-way Anova was used for furin enzymatic activity and relative wound densities with Tukey’s multiple comparison test. Western-blot quantifications were normalized by the non-treated condition and are presented as a fold change. Data are presented as means + /− SEM (standard error of the mean). P-values below 0.05 were considered statistically significant.















Leave a Reply