Functional assessment of ISG–virus interactions
We established a live-cell imaging screening system to study the virus-modulating activities of ISGs in A549 cells, which are widely used in virus research and are well-studied for their responsiveness to type-I IFNs23. The cells were engineered to stably express eGFP or mRFP-tagged histone H2B to enable a fluorescence-based read-out as a proxy for the cell number (Fig. 1a). We selected eight different fluorophore-expressing viruses belonging to different virus classes: Herpes simplex virus 1 (HSV-1) strain 17 + , Measles virus (MeV) strain EdTag, Rift Valley Fever Virus lacking Non-structural protein S (RVFVΔNSs) ZH548 strain, Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) MUC-IMB-1 strain, Semliki Forest virus (SFV) SFV6 strain, Vaccinia virus (VACV) WR strain (V300), Vesicular stomatitis virus (VSV) Indiana strain, and Yellow Fever virus (YFV) 17D strain and determined the half-maximal effective concentration (ED50) of IFNα for each virus (Supplementary Fig. 1a).

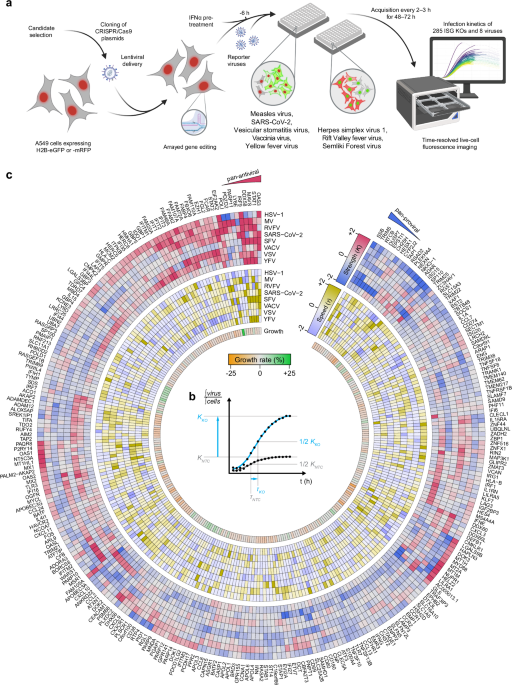
a Schematic representation of the experimental setup. A549 cells stably expressing either H2B-mRFP or H2B-eGFP were targeted for the depletion of 300 ISGs using three CRISPR/Cas9 gRNA sequence templates by lentiviral delivery. Knockout (KO) cells showing a reduction in cell growth of more than 25% compared to non-targeting control (NTC) cells were excluded. The remaining 285 ISG KO cells were pre-treated with IFNα at a virus-specific dose for 6 h and infected with eight reporter viruses. Each virus expresses a fluorophore complementary to the color of the H2B fluorophore in the cells (green virus-expressed fluorophore on H2B-mRFP and vice versa). The progression of the viral infection was captured using live-cell fluorescence imaging. Created in BioRender. Pichlmair, A. (2026) https://BioRender.com/wp5zo4e. b The ratio of the integrated virus-expressed fluorophore signal to the integrated H2B-fluorophore signal provides a normalized measure of virus infection dynamics relative to the cell number. This ratio was fitted to a logistic growth curve, allowing the quantification of key infection dynamics parameters: infection strength (K-value, upper asymptote) and the time to reach K/2 (τ-value). c Hierarchical clustering based on K-values showing the modulation of virus infection for each ISG depletion. K-values (red-to-blue), τ-values (yellow-to-blue), and the growth rate of uninfected cells (green-to-orange) are shown. All values are normalized to their respective NTC values and z-scored per virus. Shown is the mean of n = 3 biological independent replicates. Values are capped from |z | > 2.
Based on data from the Interferome database24, we selected 300 genes, of which 232 were shown to be up- (ISG) and 68 down-regulated (IFN-repressed gene, IRG) in response to type I IFNs (Supplementary Data 1). Henceforth, “ISG” will denote both ISGs and IRGs. We then stably depleted these ISGs in A549 cells using CRISPR/Cas9-based lentiviral vector pools (Fig. 1a, Supplementary Fig. 1b, c), where each gene was targeted by three sgRNA sequences selected for high on-target and low off-target scores, as well as low self-complementarity. Knockout (KO) efficiency was tested for a subset of ISGs using RT-qPCR. All tested KO cells showed significant transcript reductions for the targeted ISGs (Supplementary Fig. 1d). To avoid experimental bias, we excluded 15 ISG KO cells, which showed reduced cell growth by more than 25% in an observation period of 72 h (Supplementary Fig. 1e, Supplementary Data 2). To study the specific effect of the remaining 285 ISGs on virus infection dynamics, we induced an ISG response by pre-stimulating the KO and non-targeting control (NTC) cell lines with IFNα for 6 h prior to infection. All cell lines were then infected with the selected reporter viruses, and fluorescence images were acquired in 2–3 h intervals for 48–72 h (Fig. 1a). In total, we generated 730,802 epifluorescence images capturing the virus infection kinetics in relation to the depletion of individual ISGs. We calculated the ratio of the virus-expressed fluorophore signal and the H2B-fluorophore signal, providing a normalized measure of virus-associated reporter signal relative to the cell number (Supplementary Fig. 2a, b). We fitted this ratio to a logistic growth curve and normalized all curve parameters to the corresponding parameters of the NTC samples. The parameters describing a sigmoidal curve are the time-independent upper asymptote (K-value; representing the maximum signal reached during the observation period), the time until K/2 was reached (τ-value; representing time post infection until the half maximal signal is reached), and the slope at t = τ (β-value; representing the increase of virus growth at one timepoint) (Fig. 1b, Supplementary Data 2). Expression of H2B-fluorophore data indicated relative homogeneous expression, ensuring that the normalization of the virus-associated reporter signal to the cell number does not bias this analysis (Supplementary Fig. 2c). To accommodate the varying dynamic ranges of the reporter viruses, we employed z-scoring to normalize the parameters within each virus dataset. The parameters of viral replication curves for the NTCs closely aligned with the mean of all tested ISGs for most viruses, except for HSV-1, for which the NTC curves consistently showed reduced values for an unknown reason (Supplementary Fig. 2d, e).
To comprehensively illustrate the resulting data, we considered K– and τ-values, as well as the cell growth rate, for all targeted ISGs and viruses. We plotted these values on a heatmap using hierarchical clustering of K-values to illustrate the complex and nuanced activity of ISGs and to reveal pan-proviral and pan-antiviral activities, as well as virus-specific activities of individual ISGs (Fig. 1c, Supplementary Data 2). The IFNα signaling components STAT1, IRF9, and the translation regulator EIF2AK2 (PKR) are well-studied for their antiviral function against diverse viruses as well as their ability to regulate innate immune signaling1,7,8 and were among the ISGs with broad antiviral activities. Moreover, our data confirmed the broad antiviral activity of OAS3, a main activator of the OAS/RNase L signaling pathway25. Importantly, our analysis also correctly identified ISGs with broad antiviral functionality, such as DDX58, which predominantly exhibited antiviral activities against RIG-I-dependent RNA viruses26, while it did not show an effect for HSV-1 or SARS-CoV-2. Our data also confirmed the antiviral activity of IFI16 against HSV-1 and VACV but, in addition, highlighted its role in SARS-CoV-2 replication, which is in line with recent reports indicating that IFI16 plays a broad role in antiviral immunity for some viruses27,28,29. Notably, ISGs that are known to be functionally connected, i.e., by operating in similar pathways (e.g., STAT1 and IRF97,8) or co-dependencies (e.g., IFIT1, −2, −3—which form hetero complexes to exhibit their full activity30) clustered together in this analysis, further indicating the validity of this approach and the value of this dataset (Fig. 1c). The identification of known activities of well-characterized ISGs supports the correct and specific CRISPR/Cas9-mediated targeting and the validity to assess virus growth used in this study.
Some ISGs displayed surprisingly broad antiviral activities. These included FBXO6, an E3 ligase involved in ER-associated protein degradation (ERAD)31. FBXO6 has previously been shown to regulate immune responses and cell death induction by influenza A virus through degradation of NLRX132 and IFN type-I-mediated antiviral immunity induced by VSV, Sendai virus and synthetic ligands33. However, its broad antiviral activity has not yet been described. Furthermore, we identified an unexpected broad antiviral activity of FAM105A (OTULINL), a pseudo-deubiquitinase lacking deubiquitinase activity with a yet undescribed function34. Notably, while FAM105A has broad antiviral activity against HSV-1, MeV, RVFV, and VACV, it was proviral for VSV. Moreover, FAM20A, a poorly defined pseudokinase, exhibited similarly unexpected broad antiviral activity. Proteins linked to transcription, like the histone-lysine N-methyltransferase EZH1, as well as FAM46A (TENT5A)—a cytoplasmic non-canonical poly(A) RNA polymerase—also showed antiviral activity against multiple viruses. For FAM46A, we could also recapitulate previously reported inhibitory activity for MeV and VACV14. Similarly, FAM72A, a protein potentially involved in regulating cellular reactive oxygen species metabolism and cell growth35, was active against 5 of the 8 tested viruses.
Besides antiviral ISGs, we could identify several ISGs with proviral activity across different viruses. These ISGs included SSB, TRIM5, and RTCB. TRIM5—an E3 ubiquitin ligase that blocks reverse transcription of human immunodeficiency virus-1 (HIV-1)36,37—exhibited particularly prominent proviral activity for VACV and VSV within this dataset. The RNA chaperone La/SSB (SSB) has previously been shown to bind an internal ribosome entry site (IRES) of the Hepatitis C virus (HCV), a key component of virus translation, thereby limiting viral replication38,39. Surprisingly, our data suggests that, besides inhibiting HCV, SSB also exhibits proviral activities for HSV-1, RVFV, VACV, and VSV, indicating additional roles of SSB beyond regulating IRES-dependent translation. A similar dichotomy in virus-specific effects was observed for several other ISGs, including POLI, FAM105A, MSR1, and ZC3HAV1. Polymerase iota (POLI), involved in DNA repair40, restricted MeV and RVFV, but was required for the growth of HSV-1 and VSV. FAM105A showed antiviral effects against HSV-1, MeV, RVFV, and VACV, but was required for VSV. Similarly, the macrophage scavenger receptor 1 (MSR1) exhibited antiviral activity against SARS-CoV-2 and VACV but was found to be proviral for MeV. Another example is ZC3HAV1, which was proviral for VACV but restricted SFV and VSV replication. ZC3HAV1 has previously been shown to be a restriction factor for modified vaccinia virus Ankara (MVA)—an attenuated strain of VACV—but its depletion had no effect on VACV propagation41. Since our screen suggests that ZC3HAV1 serves as a prominent host factor for VACV propagation, an additional in-depth evaluation of ZC3HAV1’s activity in the context of poxviruses may be necessary.
The antiviral activities of ISGs were systematically tested using gain-of-function approaches13,14,16,42,43,44. Our selection of ISGs contained 111 ISGs, which were also evaluated in previous overexpression screens13,14,45. We intersected our loss-of-function data with gain-of-function studies for MeV, VACV, and YFV (Supplementary Fig. 3a). Notably, we observed that the gain-of-function screens identified a different subset of ISGs with antiviral activity, as compared to ISGs identified by the loss-of-function approach used here. The depletion of OAS3, for instance, enhanced the propagation of VACV, its overexpression had a negligible effect. Similarly, the overexpression of IRF1 robustly inhibited the replication of YFV, while the KO only had a minor effect. ISG overexpression and depletion thus serve as complementary methods for investigating cellular mechanisms, illustrating the added value of both strategies.
To evaluate these findings in different cell types, we depleted a subset of selected ISGs in SK-N-SH neuroblastoma cells and primary human foreskin fibroblasts (HFF) and challenged them with RVFV, HSV-1, and VACV (Supplementary Fig. 3b, c). These experiments showed that the majority of ISG functions were conserved between different cell types, but we could also identify virus- and cell-type-specific activities. For HSV-1, for instance, antiviral functions were similar in all tested cell lines, but the magnitude of the response was generally lower in SK-N-SH cells and HFFs. Some individual ISGs appear to differ in their antiviral properties among the tested cell types. The antiviral effect of NUPR1 against VACV, for instance, was particularly prominent in HFFs as compared to other tested cell types in this screen. Notably, some ISGs had cell-type-specific functions. An example was HELZ, which was proviral for VACV in A549 cells but antiviral in HFFs and SK-N-SH cells. Similarly, DDX60 and MX2 had opposing effects in RVFV-infected A549 and HFFs. This analysis shows that the majority of ISGs had similar activities in different cell types, but that a subset of ISGs have more pronounced or nuanced effects in specific cell types, which is in line with other reports on cell-type specificity of ISGs1,46,47,48,49.
Collectively, our data revealed pro- and antiviral properties of a large number of ISGs with yet unreported functions. While a relatively small subset of ISGs was broadly active against many viruses, we identified numerous ISGs that showed virus-specific effects.
Delineating ISGs’ specificity to modulate virus growth
The depletion of most ISGs had minimal or no impact on most tested viruses (Fig. 2a). A few ISGs per virus displayed major effects (|z| ≥ 2 standard deviations (SDs)), while many ISGs appeared to only subtly regulate virus infection (0.5 ≤ |z | < 2 SDs). Approximately one quarter of the tested ISGs displayed pro- and antiviral activity against individual viruses. This suggests that the combined activities of multiple ISGs contribute to the total antiviral activity against individual viruses. The most robust antiviral activities per virus were observed for EIF2AK2 (HSV-1), PARP11 (MeV), IRF9 (RVFV and YFV), LY6E (SARS-CoV-2), OAS3 (VACV), and IFITM1 (VSV) (Fig. 2b). Lymphocyte antigen 6E (LY6E) was identified to be active against SARS-CoV-2, which aligns with its known role in restricting SARS-CoV-2 entry17,50. Moreover, correctly identifying well-studied ISGs such as IRF9, EIF2AK2, OAS3, and IFITM1 among the most prominent hits1 validated our screening approach. This analysis also identified several ISGs with yet unreported antiviral activity, like POLI and PDZD2—a protein similar to pro-interleukin-1651—which showed antiviral activity against MeV. The same analysis also allowed us to identify ISGs with specific virus-promoting functions. These included LGALS9B for MeV and RVFV, ATXN1 for SARS-CoV-2, ADAMDEC1 for SFV, RTCB for VSV, GPR141 for YFV, POLI for HSV-1 and ZC3HAV1 for VACV.

a Categorical effects of ISG knockouts (KOs) based on their K-values: neutral (gray), minor pro- or antiviral (0.5 < |z | ≤ 2 SDs, light blue and light red, respectively), and major pro/antiviral (|z | ≥ 2 SDs, blue and red, respectively). SD: standard deviation. b Beeswarm plots of K–values after z-scoring, showing all assessed KO effects. c Concatenated ISG effects with respect to K-values across all viruses, categorizing ISG KOs as virus-specific, pan-antiviral, pan-proviral, or neutral. ISGs showing major effects against a single virus but only minimal or no effects for the other viruses were considered virus-specific. ISGs exhibiting pro- or antiviral effects among at least six of the eight tested viruses were considered pan-proviral or pan-antiviral, respectively.
For τ-values (time to reach K/2), we observed a similarly large subset of ISGs altering the infection progression (Fig. 1c, Supplementary Fig. 3d). Among these, the candidates causing the most pronounced delay in half maximum infection signal were RIN2 (HSV-1), PARP11 (MeV), STAT2 (RVFV), LY6E (SARS-CoV-2), OAS3 (SFV and VACV), OAS1 (VSV), and IRF9 (YFV) (Fig. 1c, Supplementary Fig. 3e). On the other hand, candidates causative for an accelerated time to K/2 were POLI (HSV-1), HELZ2 (MeV), MAVS (RVFV), CXCL11 (SARS-CoV-2), EIF2AK2 (SFV), ZC3HAV1 (VACV), IFI16 (VSV), and IFI35 (YFV).
To concatenate the data more comprehensively, we combined K-values from all viruses to identify ISGs with consistent pro- or antiviral effects among at least six of the eight viruses tested (Fig. 2c). When considering both minor and major effects, five candidates—FAM110A, FCAR, FGL2, IRF9, and STAT1—were broadly antiviral. At the same time, APOBEC3A, RTCB, SSB, and TRIM5 were pan-proviral. This multi-virus analysis revealed key ISGs, such as LY6E and POLI, with solid effects on the replication of specific viruses while highlighting novel pan-anti and proviral ISGs.
A machine-learning approach to identify functional relationships
To further investigate the complex relationship between ISGs and the tested viruses, we employed an unsupervised machine-learning approach. Self-organizing maps (SOM) iteratively learn the underlying data structure in a multi-dimensional space, represented by the z-scored input parameters—K-, τ-, and β-values for all candidates and viruses—and assign a categorical neuron to each ISG (Fig. 3a, Supplementary Fig. 4a). Because the SOM is trained on z-scored parameters, it primarily captures relative patterns and correlations across viruses rather than absolute effect magnitudes. The distribution of ISGs across the SOM revealed intriguing patterns of virus-host interactions (Fig. 3b). For instance, some neurons contained many ISGs with only minor effects on viral infection (Fig. 3b; e.g., neurons 11, 16), while other neurons contained few ISGs with dominant effects (Fig. 3b; neurons 2, 3, 5, 6, 12, 13, 17, 20). Notably, many ISGs with similar functions were grouped into the same neuron. This is exemplified by proteins involved in type-I IFN signaling, such as DDX58, IRF9, MAVS, STAT1, and STAT2, all assigned to neuron 5. This approach also revealed some unexpected similarities between ISGs. In neuron 20, only two members were assigned, POLI and PARP11. As both are involved in DNA damage repair40,52 and appear to have a similar impact across different virus infections, our analysis might have identified a close functional relationship between these proteins in the context of virus infections. Furthermore, TRIM25 and ZC3HAV1 are the sole members of neuron 17. TRIM25 plays a crucial role in the early stages of the immune response to viral infections, primarily by activating the RIG-I pathway53. ZC3HAV1, also known as zinc-finger antiviral protein (ZAP), is a well-described restriction factor for several viruses through targeting viral RNAs54,55,56,57. TRIM25 has previously been shown to be a cofactor of ZC3HAV158,59,60, thus contextualizing the functional clustering of these two proteins into the same neuron. Additionally, the SOM identified ISGs with distinct activities. More specifically, LY6E, OAS3, RNA 2’,3’-cyclic phosphate and 5’-OH ligase (RTCB), and EIF2AK2 were placed in distinct positions in the SOM and are the sole members of neurons 2, 3, 6, and 13, respectively. RTCB, an atypical RNA ligase and integral core component of the tRNA-splicing ligase complex (tRNA-LC), influenced the infection dynamics of multiple viruses, including HSV-1, MeV, RVFV, SARS-CoV-2, VACV, and VSV (Fig. 3c). Importantly, the depletion of RTCB did not significantly affect cell growth within 72 h of observation (Supplementary Fig. 4b). These results prompted us to validate the activity of RTCB. To understand the molecular mechanisms underlying the proviral activity of RTCB, we employed quantitative proteome expression analysis. We pre-treated RTCB KO and NTC cells with 500 U/mL IFNα for 16 h to induce an ISG response and then used liquid chromatography coupled with tandem mass spectrometry (LC-MS/MS) to analyze proteomic changes. Notably, this analysis revealed a significant downregulation of all subunits of the tRNA-LC (C14orf166, C2orf49, DDX1, FAM98B, and RTCB) in RTCB KO cells61 (Fig. 3d, Supplementary Fig. 4c, Supplementary Data 3). Functional enrichment analysis based on CORUM terms62 revealed effects on the tRNA-LC, as well as terms related to the Spliceosome, the pre-rRNA complex, ribosomal subunits, and exosomes (Supplementary Fig. 4d, Supplementary Data 3). We expressed V5-tagged RTCB and control proteins in A549 cells and analyzed α-V5-precipitated proteins using LC-MS/MS. Among other proteins, the protein–protein interactome identified all five components of the tRNA-LC as significant interactors of RTCB (Supplementary Fig. 4e, Supplementary Data 4). Enriched CORUM terms of the interactors showed associations to chaperone-related complexes like the CCT complex, the BBS-chaperonin complex, and the CALM1-FKBP38-BCL2 complex (Supplementary Fig. 4f, Supplementary Data 4). This combined analysis suggests that RTCB binds to and stabilizes the tRNA-LC and its associated functions, while potentially being involved in other pathways, such as RNA processing and protein folding.

a Self-Organizing Maps (SOMs) project high-dimensional data onto a lower-dimensional grid of neurons. During training, the input data (K–, τ-, and β-values across all eight viruses and 285 ISG KOs) are clustered to represent, as faithfully as possible, the similar localization of the data points in the multi-dimensional space into the grid of neurons. Each KO datapoint is then associated with the closest neuron. b SOM with a total of 20 neurons. The background color indicates the number of ISG KOs assigned to a particular neuron. The line plot within each neuron shows the codebook of that neuron, reflecting the weight vector derived from the input data. A high-weight vector value indicates that the input parameters strongly influence the response of the neuron, suggesting a pronounced effect of the corresponding ISG on viral infection kinetics. Neurons with pronounced effects are further detailed, displaying their members surrounding the map. c Virus-derived fluorophore signals normalized to the H2B-mRFP or -eGFP signal measured by live-cell imaging. The sigmoidal curves, calculated from the mean parameters of all replicates, compare RTCB KO (cyan) with the average of five non-targeting control (NTC) samples (gray) for each virus. d Box plots showing iBAQ intensities of tRNA ligase complex (tRNA-LC) subunits in RTCB KO cells as determined by proteomic analysis. Box plots show the median (center line), first and third quartiles (box limits), and whiskers extending up to 1.5× the interquartile range. A two-tailed, two-sample Welch’s t-test was performed (n = 4 biologically independent samples), followed by a false discovery rate (FDR) correction of p-values.
In conclusion, the SOM approach provides a comprehensive overview of the complex activity of ISGs in viral infections and offers a predictive tool for future research. ISGs clustering to similar neurons may share similar properties, as illustrated by the grouping of functionally related ISGs, such as STAT1, STAT2, IRF9, and PRRs, that stimulate IFN induction and TRIM25 and ZC3HAV1, which are functionally connected. Moreover, the SOM allowed the identification of ISGs with unique functions, such as RTCB.
Analysis of ISG activities against SARS-CoV-2
Regularly occurring viral epidemics and pandemics necessitate a deeper understanding of the functional role of host factors, which have the ability to control these viruses. To visualize the impact of all ISGs on SARS-CoV-2, we plotted K-value (max fluorophore expression value) and τ-value (time to reach K/2) obtained from all tested KO cells (Fig. 4a, b). As expected17,50, the depletion of LY6E prominently increased the K-value. Similarly, our data recapitulated STAT2 and IRF9 as antiviral factors. Several other ISG KOs demonstrated pronounced antiviral activity against SARS-CoV-2, such as PARP10 and CFB. PARP proteins were shown to have antiviral activity against Alphaviruses63,64,65, while deregulation of the complement system involving CFB was shown to play an important role in Dengue virus infection66,67. However, an antiviral activity against coronavirus has not been reported so far. Further unexpected findings were the antiviral activities of Tryptophanyl-tRNA Synthetase 1 (WARS1), which is essential to attach tryptophan to its cognate tRNA, and BORC subunit 8 (BORCS8, MEF2BNB), a protein involved in cellular trafficking and organelle biogenesis68. We also identified ISGs, which, when depleted, reduce virus infection dynamics. Among them was LGALS3BP, a protein reported as a negative regulator of the IFN response45,69, which may explain the reduced SARS-CoV-2 infection observed upon its depletion. A similar reduction of SARS-CoV-2 propagation was observed by depletion of TDRD7, a protein implicated in stress granule formation, translational silencing, and autophagy70. TDRD7 is a restriction factor for VACV and VSV71,72, which we could recapitulate in our screen (Fig. 1c). Unexpectedly, we could identify proviral activities for ATXN1, a protein involved in transcriptional repression and RNA splicing73, and SAMHD1, best described for its antiviral activity against HIV-1 and other retroviruses by inhibiting viral DNA synthesis69.

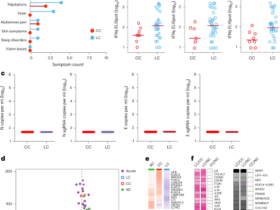
a Scatter plot showing normalized infection strength (K-value) and speed (τ-value) for all ISG knockout (KO) cells infected with SARS-CoV-2. Triangles denote ISGs selected for follow-up experiments. b Table showing selected ISGs along with their respective K-, τ-, β-values, and growth rates; color gradients correspond to the column-specific minimum and maximum values. c Schematic of experimental design used for proteomic analysis of ISG KO and ISG overexpressing (OE) cells, loss-of-function (LOF), and gain-of-function (GOF), respectively. KO cells were pre-treated with IFNα (6 h, 6.25 U/mL), infected with SARS-CoV-2 or left uninfected for 24 h and subjected to proteome analysis. Cells overexpressing V5-tagged ISGs were pre-treated with 500 U/mL IFNα for 16 h. All samples were subjected to proteome and interactome analysis using α-V5 affinity purification. The resulting proteome data were analyzed for enriched Gene Ontology Biological Process (GO:BP) terms. Intersections of terms identified in the interactome data and at least two other datasets (cyan intersects) were then used to describe the function of the ISG. Created in BioRender. Pichlmair, A. (2026) https://BioRender.com/3op6pws. d UpSet plots showing the intersection of enriched GO:BP terms derived from the significantly enriched proteins of the datasets depicted in c. The bar plots on the left of each UpSet plot show the total number of enriched GO:BP terms per dataset. The number of shared terms among datasets is shown at the top of each intersection. The terms present in at least two proteome datasets and the interactome dataset (cyan intersects) were selected and are shown below each plot. The terms were manually filtered for redundancy and generic terms.
We reasoned that ISGs with pro- or antiviral functions are involved in pathways that are particularly important for SARS-CoV-2 replication. To identify such pathways, we performed orthogonal proteomic profiling of cells that lack or overexpress the 19 ISGs with the most prominent SARS-CoV-2 phenotypes (Fig. 4c). To determine the cellular interaction partners for the selected ISGs, we expressed the 19 candidates as V5-tagged proteins in A549 cells (Fig. 4b, c). Cells were stimulated with IFNα (500 U/mL, 16 h) to mimic the proteome composition of an antiviral response and subsequently used for affinity purification using α-V5 beads followed by LC-MS/MS analysis. We confirmed the successful identification of all bait proteins except for LY6E, which was not detected, and identified 10–40 interactors per bait (Supplementary Fig. 5a, b, Supplementary Data 5). Overlap with known interactors of SARS-CoV-274 with ISG interactors revealed 32 overlapping proteins in both datasets (Supplementary Fig. 5c), which could serve critical roles in mediating anti-SARS-CoV-2 activity. We further determined the differential protein expression pattern of A549 cells overexpressing the 19 ISGs (Fig. 4c), which reveals pathways that are regulated by the selected ISGs and mimics gain-of-function experiments (Supplementary Data 5). These data included 329 known SARS-CoV-2 interactors which could be relevant to mediate antiviral activity against SARS-CoV-2 (Supplementary Fig. 5d). We also characterized protein expression in cells depleted for the individual ISGs and pre-treated with IFNα (6.25 U/mL for 6 h) or pre-treated with IFNα followed by SARS-CoV-2 infection, respectively. We confirmed a significant depletion of eight ISGs, which were detectable by LC-MS/MS (Supplementary Fig. 5e). Among the 6,379 quantified proteins, we identified 50–150 proteins that were significantly regulated in each of the individual ISG KO cells (Supplementary Fig. 5f, Supplementary Data 6). As expected, depletion of STAT2 and IRF9 led to an impaired antiviral innate immune response pattern (e.g., reduced EIF2AK2, IFIT1, ISG15, OAS3, and STAT1) (Supplementary Fig. 5g). To concatenate these datasets and to filter for the dominant pathways regulated by the selected ISGs, we intersected GO:BP terms that were enriched in the interactome and at least two protein expression datasets (Fig. 4d, Supplementary Fig. 6a, Supplementary Data 7). This strategy successfully recapitulated the function of STAT2 as a key protein in the innate immune response, as indicated by the associated terms “type I interferon-mediated signaling” and “defense response to virus”, among others. FBXO6 exhibited terms associated with ubiquitination, proteasomal degradation, and protein folding, aligning with its known function in endoplasmic reticulum-associated degradation31,75. Interestingly, the ubiquitin-dependent degradation of proteins associated with FBXO6 and its potential involvement in protein dynamics during viral replication may explain the broad antiviral activity of FBXO6 (Fig. 1c, Fig. 4d). Similarly, the term “organelle localization” for BORCS8 matched its known function in lysosome localization and motility. Enrichment of other terms, such as RNA processing and translation regulation, suggests a multifaceted role of BORCS8 in controlling viral replication. Moreover, WARS1 was linked to RNA processing and translation, among other functions, which are also congruent with its role in protein synthesis.
Our analysis revealed a diverse range of cellular processes that contribute to the inhibition of SARS-CoV-2. Among the processes most significantly affecting SARS-CoV-2 growth are IFN signaling, protein stability and degradation, RNA processing, transcription and translation, and trafficking (Fig. 4d, Supplementary Fig. 6a). This suggests that targeting proteins or a combination of proteins involved in these processes would be valuable for perturbing virus infection. Moreover, we confirmed the antiviral activities of key ISGs such as LY6E, STAT2, and IRF9.
Synergistic activity of ISGs
Since ISGs are co-expressed after exposure to type-I IFN, we tested whether the combined depletion of any of the 19 selected ISGs may potentiate their effect on SARS-CoV-2 propagation. We thus depleted the selected ISGs in all possible pairwise combinations and measured the progression of SARS-CoV-2-GFP replication by fluorescence live-cell imaging (Fig. 5a). We assumed a multiplicative relationship between the single KO effects and thus calculated the difference between the product of the two single effects to the experimentally evaluated effect of ISG co-depletion (Fig. 5b). The resulting interaction scores were then plotted in a heatmap using hierarchical clustering (Fig. 5c, Supplementary Data 8). Co-depletion of the majority of tested ISGs equaled the expectations considering the product of two individual KOs. The co-depletion of LY6E, BORCS8, and STAT2 with other candidates had particularly beneficial effects that exceeded the expected effects of the individual KOs. This was most prominent for the co-depletion of LY6E with either STAT2 or IRF9. These results indicate that concurrent activation of IFN signaling through STAT2 and inhibition of viral entry through LY6E exerts a synergistic antiviral effect against SARS-CoV-2. Similar synergisms were observed for the depletion of BORCS8 with STAT2 and IRF9, respectively. However, the co-depletion of some ISGs also resulted in lower-than-expected values, as indicated by negative interaction scores, suggesting that depleting both proteins did not confer any additional impact. This was particularly evident for the co-depletion of STAT2 and IRF9 and several other ISG combinations (ATXN1 with VCAN, ATXN1 with OAS1, OAS1 with WARS1, SAMHD1 with PARP10, and LY6E with BORCS8) (Fig. 5c). Plausible explanations for this phenotype may be that the respective activity of these ISGs may depend on each other or contribute to similar cellular functions. Indeed, STAT2 and IRF9 contribute to the same pathway (IFN signaling), which explains why their simultaneous depletion does not lead to a greater effect compared to single depletions. A similar relationship could be envisioned for, e.g., SAMHD1, which regulates the intracellular nucleotide pool, and PARP10, which requires nucleotides for poly-ADP ribosylation. Interestingly, the co-depletion of BORCS8 and LY6E did not lead to the expected increase in antiviral activity. This led us to hypothesize that BORCS8 and LY6E may operate in a similar pathway or inhibit the same stage of the viral life cycle. Since LY6E inhibits virus entry17,50, we envisioned that BORCS8 may be similarly involved in SARS-CoV-2 infection. To test this, we employed GFP-encoding VSV particles that were pseudotyped with the SARS-CoV-2 Spike protein (VSV-S), thereby utilizing the SARS-CoV-2 entry route into host cells. Depletion of BORCS8 significantly enhanced infection by VSV-S but not of VSV particles bearing their natural glycoprotein (G) (Fig. 5d), supporting the involvement of BORCS8 in SARS-CoV-2 entry.

a Diagram illustrating the co-depletion of ISGs and evaluation of their impact on SARS-CoV-2-GFP infection. ISG knockouts (KOs) were performed sequentially by transducing cells with CRISPR/Cas9 lentiviral vectors encoding puromycin or hygromycin resistance. After selection, the co-depleted KO cells were infected with SARS-CoV-2-GFP (MOI 3). Fluorescence images were acquired using live-cell fluorescence imaging every 4 h for 72 h. Created in BioRender. Pichlmair, A. (2026) https://BioRender.com/fgxwlao. b Schematic to evaluate interaction scores. For each replicate, K-values were normalized to the median value across all co-depleted KO cells. K-values of ISG KOs paired with an NTC were considered as single KO effects. We estimated the anticipated double KO (dKO) effect, by multiplying these individual effects. Subsequently, the interaction score was calculated by subtracting the estimated dKO effect from the observed dKO effect. A positive interaction score indicates a synergistic effect beyond the expected combined individual effects, while a negative score suggests an antagonistic interaction. c Interaction scores of all dKO combinations, displayed following hierarchical clustering. Redundant combinations represent duplicated values. Shown is the mean (n = 4 biologically independent samples). d GFP-expressing VSV-Spike pseudotyped virus infection compared to VSV-G in BORCS8 KO, LY6E KO, and NTC cells. All data are normalized to the corresponding NTC cells, as measured by live-cell fluorescence imaging. Peak signal intensity was observed at 24 h for VSV-G and 48 h for VSV-Spike. Data are presented as mean ± SD. One-sided, two-sample Welch’s t-test (n = 3 biologically independent experiments).
Through this systematic analysis, we demonstrated that co-depletion of specific antiviral ISGs, such as LY6E with STAT2 or IRF9, prominently supports SARS-CoV-2 growth, indicating potent synergistic activity against SARS-CoV-2 when concomitantly disrupting viral entry and stimulating IFN signaling.
BORCS8 impacts vesicle trafficking and endosomal dynamics
We decided to further evaluate the role of BORCS8 on SARS-CoV-2 infection. The effect of depleting BORCS8 showed a prominent effect for SARS-CoV-2 in which the KO led to increased virus infection. We also observed moderate antiviral effects for HSV-1 and YFV (Fig. 1c, Supplementary Fig. 7a). Analysis of the genomic locus by Sanger sequencing confirmed correct targeting of BORCS8 (Supplementary Fig. 7b) and the depletion of BORCS8 did not affect growth rates as compared to NTC cells (Supplementary Fig. 7c). In line with the screening data, SARS-CoV-2 infected BORCS8 KO cells accumulated significantly more infectious virus particles in their supernatants for the parental strain and variant of concern (VOC) B.1.617.2 (Delta variant) of SARS-CoV-2, as compared to infected control cells (Fig. 6a). Similarly, BORCS8 depletion yielded significantly higher viral RNA accumulation of VOCs B.1.617.2 (Delta variant) and BA.1 (Omicron variant) as measured by RT-qPCR (Supplementary Fig. 7d). Compared to NTC cells, CRISPR/Cas9-mediated depletion of BORCS8 in Calu-3 cells also led to an increase in SARS-CoV-2 growth, which was comparable to the depletion of LY6E (Fig. 6b), suggesting that BORCS8 exerts antiviral activity across cell types. Expression of CRISPR/Cas9-resistant BORCS8 in BORCS8 KO cells reversed the enhanced viral replication, confirming the specificity of the KO approach (Fig. 6c).

a Plaque assay comparing SARS-CoV-2 accumulation in BORCS8 knockout (KO) and NTC supernatants infected with the parental strain and B.1.617.2 (Delta) variant of SARS-CoV-2 (MOI 3) at 24 h post-infection. Shown is the mean ± SD (n = 3 biologically independent experiments). One-sided, two-sample Welch’s t-test. b Calu-3 cells were infected with SARS-CoV-2-GFP (MOI 3), and the GFP signal was measured by live-cell imaging. The figure shows the integrated intensity of the virus-expressed GFP signal normalized to the confluence of the cells at 60 h post-infection. Data are presented as mean ± SD (n = 3 biologically independent experiments). One-sided, two-sample Welch’s t-test. c A549-ACE2 cells of the indicated genotype were infected with SARS-CoV-2-GFP (MOI 3) and imaged at 2 days post-infection. GFP intensity was normalized to NTC. Mean ± SD (n = 3 biologically independent experiments). One-sided, one-sample t-test. d Representative Airyscan images of NTC and BORCS8 KO cells stained for LAMP1, α-Tubulin, and DAPI. Maximum intensity projection. Scale bar: 25 µm. e Effect of TMPRSS2 expression and Camostat treatment on SARS-CoV-2 N RNA levels in BORCS8 KO and NTC A549-ACE2 cells at 24 h post-infection. Cells were treated with 10 µM Camostat for 6 h prior to the infection. SARS-CoV-2 N RNA was quantified by RT-qPCR and normalized to the mean of NTC samples. Bars show mean ± SD of –ΔΔCt values (n = 3 biologically independent experiments). One-way ANOVA, followed by Tukey’s HSD post hoc test for multiple comparisons. SD standard deviation.
The depletion of BORC subunits has been shown to result in a dysfunctional BORC complex, leading to impaired lysosomal motility, which can be characterized by the perinuclear localization of LAMP1-stained vesicles68,76. Indeed, Airyscan microscopy of LAMP1 revealed a pronounced perinuclear accumulation of lysosomes in BORCS8 KO cells, confirming functional targeting of the BORC complex (Fig. 6d). Lysosomes are known to be relevant for SARS-CoV-2 entry and egress. We hypothesized that a potential inhibitory effect of BORCS8 on endosomal uptake of SARS-CoV-2 can be alleviated by facilitating plasma membrane fusion of SARS-CoV-2 by expressing TMPRSS2 in BORCS8 KO cells. As expected, TMPRSS2 expression increased viral RNA levels in NTC cells, indicating enhanced plasma membrane entry (Fig. 6e). The TMPRSS2 inhibitor camostat reversed this effect, confirming specificity. Notably, expression of TMPRSS2 or camostat treatment had no effect in BORCS8 KO cells, indicating that BORCS8 controls the endosomal uptake route of SARS-CoV-2.
Previously, ORF3a of SARS-CoV-2 was shown to associate with lysosomes to exploit lysosomal exocytosis for viral egress76,77,78. The fusion of lysosomes with the plasma membrane releases nascent virus particles into the extracellular space while simultaneously transferring lysosome-associated proteins onto the plasma membrane. Since BORCS8 depletion affected the lysosomal compartment (Fig. 6d), we investigated whether BORCS8 might influence the correct localization of ORF3a. As expected, exogenously expressed ORF3a localized to both the cytoplasm and the plasma membrane in control cells (Supplementary Fig. 8a). In BORCS8 KO cells, however, the localization of ORF3a was exclusively cytoplasmic, indicating defective ORF3a-trafficking due to BORCS8 depletion. A prominent role of BORCS8 in endosomal trafficking was also evident from proteome analysis of cells overexpressing or depleted for BORCS8 (Fig. 4d). Differentially expressed proteins and associated enrichment terms of IFNα pre-treated BORCS8 KO cells indicated perturbations of proteins involved in translation and in the BLOC1/2 complex, which is critical for endosomal cargo sorting, and the endosomal targeting complex (Fig. 7a). Moreover, BORCS8 KO cells infected with SARS-CoV-2 revealed perturbation of the CCC and the CCC-Wash complex (Fig. 7b), which mediate endosomal sorting, protein trafficking, and actin polymerization on endosomal membranes79. BLOCS8 co-precipitated with the majority of BORC complex subunits (BORCS5, BORCS8, C17orf59, C10orf32, KXD1, and LOH12CR1) but also components of the related BLOC complex (BLOC1S1, BLOC1S2, and SNAPIN) (Fig. 7c). The associated enrichment terms further emphasize the role of BORCS8 in the BORC and BLOC complexes, and the v-ATPase-Ragulator complex, an essential component for lysosomal function (Fig. 7d). Additionally, the interactome of BORCS8 in the IFNα-treated condition enriched for the TRBP-containing complex, which is involved in RNA silencing, and the TNF/NF-κB signaling complex 6, which is integral for inflammatory responses (Fig. 7e).

a–e A549-ACE2 cells were analyzed as shown in Fig. 4c. a-b and d-e. Gene set enrichment analysis based on the indicated proteomic dataset (4 biologically independent samples) using g:Profiler and a one-sided hypergeometric test. p-values were adjusted for multiple comparisons using the g:SCS algorithm. The most significant CORUM terms are shown. p-values exceeding the maximum range are capped and indicated with a triangle. c Volcano plot showing protein interactors detected by interactome analysis in A549 cells overexpressing BORCS8. Subunits of the BORC and BLOC complex are indicated in cyan and orange. Two-tailed, two-sample Welch’s t-test (n = 3 biologically independent samples). Protein groups with an |log2FC | ≥ 2 and a p-value ≤ 0.05 were considered significant and are indicated by a dotted line. f Time-course analysis of acidic vesicles in BORCS8 KO and NTC cells stained with LysoTracker Red. The mean ± SD is shown as line and ribbons (n = 4 biologically independent samples). The area under the curve (AUC) was calculated for each replicate and subjected to a two-tailed, two-sample Welch’s t-test. g As f), but cells were treated with pHrodo Green. h Time-resolved colocalization of endocytosed cargo (pH-insensitive dye, Dextran-AF647) with endocytosed and acidified cargo (pH-sensitive dye, pHrodo Green). Confocal images were acquired every 6.5 s for a total of 7 min. The colocalization was carried out by Otsu’s thresholding for each channel and time-point and calculating the number of double-positive pixels normalized to the number of Dextran-AF647-positive pixels (total endocytosed cargo). Individual values and the loess-smoothed trend over time are shown.
Based on the involvement of BORCS8 in endosomal cargo sorting and given the increased entry of Spseudotyped VSV particles in BORCS8-depleted cells, we specifically focused on the role of BORCS8 in endocytosis. We treated cells with LysoTracker Red, a pH-sensitive dye that stains acidic vesicles in the cell, and performed live-cell imaging of BORCS8 KO and control cells. Notably, compared to controls, BORCS8-deficient cells showed increased levels of acidified vesicles, as indicated by the integrated intensity of LysoTracker (Fig. 7f). Reconstitution of KO cells with BORCS8 reversed the increased LysoTracker signal to levels observed in control cells (Supplementary Fig. 8b). The increased acidification of endosomes could arise from malfunctions in either the maturation or fusion processes of endosomes and lysosomes, which are crucial for the degradation of endocytosed cargo and the recycling of cellular components. In line with this, we observed higher levels of vesicles labeled with pHrodo, a pH-sensitive dye that is internalized by endocytosis, allowing for the evaluation of early endosome acidification (Fig. 7g). In BORCS8-depleted cells, endocytosed cargo accumulated to higher levels, indicating retention of endocytosed cargo in the process of endosome maturation. Interestingly, when evaluating time-resolved colocalization of a pHrodo and a pH-insensitive dye Dextran Alexa Fluor 647, representative of endosome acidification and endocytosis activity, respectively, we could observe a defect in the acidification of incoming cargo in BORCS8 KO cells (Fig. 7h, Supplementary Fig. 8c). However, acidified vesicles and the contained cargo appear to not progress into more mature endo-lysosomes, which aligns with a role of BORCS8 in endocytosis and lysosomal function. Collectively, depletion of BORCS8 alters endo-lysosomal dynamics and enhances endocytic uptake, suggesting that BORCS8 constrains SARS-CoV-2 infection by maintaining proper vesicle processing and restricting entry via the endocytic pathway.












Leave a Reply