Klein, L. & Petrozziello, E. Antigen presentation for central tolerance induction. Nat. Rev. Immunol. 25, 57–72 (2025).
Wolfl, M. et al. Hepatitis C virus immune escape via exploitation of a hole in the T cell repertoire. J. Immunol. 181, 6435–6446 (2008).
ElTanbouly, M. A. & Noelle, R. J. Rethinking peripheral T cell tolerance: checkpoints across a T cell’s journey. Nat. Rev. Immunol. 21, 257–267 (2021).
Brown, C. C. & Rudensky, A. Y. Spatiotemporal regulation of peripheral T cell tolerance. Science 380, 472–478 (2023).
Vidovic, D. & Matzinger, P. Unresponsiveness to a foreign antigen can be caused by self-tolerance. Nature 336, 222–225 (1988).
Richards, D. M., Kyewski, B. & Feuerer, M. Re-examining the nature and function of self-reactive T cells. Trends Immunol. 37, 114–125 (2016).
Yu, W. et al. Clonal deletion prunes but does not eliminate self-specific alphabeta CD8+ T lymphocytes. Immunity 42, 929–941 (2015).
Danke, N. A., Koelle, D. M., Yee, C., Beheray, S. & Kwok, W. W. Autoreactive T cells in healthy individuals. J. Immunol. 172, 5967–5972 (2004).
Wortel, I. M. N., Kesmir, C., de Boer, R. J., Mandl, J. N. & Textor, J. Is T cell negative selection a learning algorithm? Cells 9, 690 (2020). This paper presents a computational model showing that the choice of thymic self peptides can optimize self–non-self discrimination, drawing parallels between TCR recognition of pMHC and language learning.
Huseby, E. S. et al. How the T cell repertoire becomes peptide and MHC specific. Cell 122, 247–260 (2005).
Chu, H. H. et al. Positive selection optimizes the number and function of MHCII-restricted CD4+ T cell clones in the naive polyclonal repertoire. Proc. Natl Acad. Sci. USA 106, 11241–11245 (2009).
Mandl, J. N., Monteiro, J. P., Vrisekoop, N. & Germain, R. N. T cell-positive selection uses self-ligand binding strength to optimize repertoire recognition of foreign antigens. Immunity 38, 263–274 (2013). This study, together with Persaud et al. (2014) and Fulton et al. (2015), demonstrates that naive CD4+ and CD8+ T cells with differences in self-reactivity differ in their function upon activation.
Fulton, R. B. et al. The TCR’s sensitivity to self peptide-MHC dictates the ability of naive CD8+ T cells to respond to foreign antigens. Nat. Immunol. 16, 107–117 (2015).
Persaud, S. P., Parker, C. R., Lo, W. L., Weber, K. S. & Allen, P. M. Intrinsic CD4+ T cell sensitivity and response to a pathogen are set and sustained by avidity for thymic and peripheral complexes of self peptide and MHC. Nat. Immunol. 15, 266–274 (2014).
Mandl, J. N., Torabi-Parizi, P. & Germain, R. N. Visualization and dynamic analysis of host-pathogen interactions. Curr. Opin. Immunol. 29, 8–15 (2014).
Shapiro, I. E. & Bassani-Sternberg, M. The impact of immunopeptidomics: from basic research to clinical implementation. Semin. Immunol. 66, 101727 (2023).
Caron, E. et al. Analysis of major histocompatibility complex (MHC) immunopeptidomes using mass spectrometry. Mol. Cell. Proteom. 14, 3105–3117 (2015).
Kubiniok, P. et al. Understanding the constitutive presentation of MHC class I immunopeptidomes in primary tissues. iScience 25, 103768 (2022).
Bozzacco, L. et al. Mass spectrometry analysis and quantitation of peptides presented on the MHC II molecules of mouse spleen dendritic cells. J. Proteome Res. 10, 5016–5030 (2011).
Ploegh, H. L., Orr, H. T. & Strominger, J. L. Major histocompatibility antigens: the human (HLA-A, -B, -C) and murine (H-2K, H-2D) class I molecules. Cell 24, 287–299 (1981).
Harding, C. V. & Unanue, E. R. Quantitation of antigen-presenting cell MHC class II/peptide complexes necessary for T-cell stimulation. Nature 346, 574–576 (1990).
Klein, L., Kyewski, B., Allen, P. M. & Hogquist, K. A. Positive and negative selection of the T cell repertoire: what thymocytes see (and don’t see). Nat. Rev. Immunol. 14, 377–391 (2014).
Takada, K., Ohigashi, I., Kasai, M., Nakase, H. & Takahama, Y. Development and function of cortical thymic epithelial cells. Curr. Top. Microbiol. Immunol. 373, 1–17 (2014).
Honey, K. & Rudensky, A. Y. Lysosomal cysteine proteases regulate antigen presentation. Nat. Rev. Immunol. 3, 472–482 (2003).
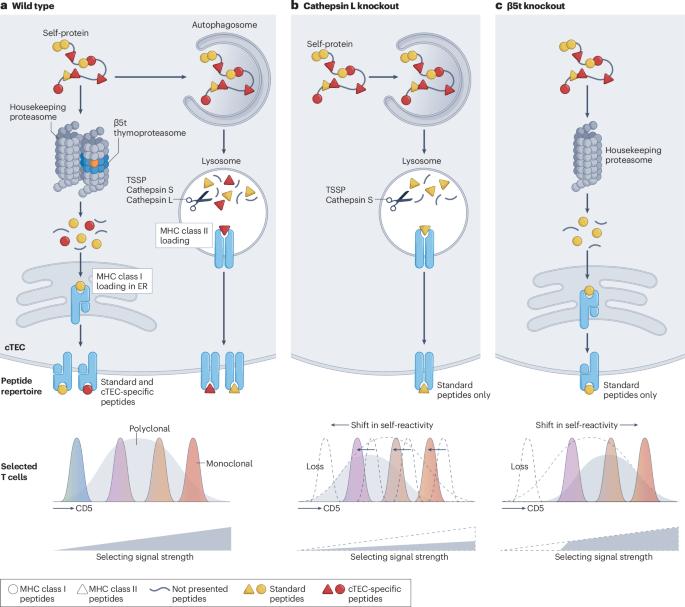
Murata, S. et al. Regulation of CD8+ T cell development by thymus-specific proteasomes. Science 316, 1349–1353 (2007).
Nakagawa, T. et al. Cathepsin L: critical role in Ii degradation and CD4 T cell selection in the thymus. Science 280, 450–453 (1998).
Honey, K., Nakagawa, T., Peters, C. & Rudensky, A. Cathepsin L regulates CD4+ T cell selection independently of its effect on invariant chain: a role in the generation of positively selecting peptide ligands. J. Exp. Med. 195, 1349–1358 (2002).
Gommeaux, J. et al. Thymus-specific serine protease regulates positive selection of a subset of CD4+ thymocytes. Eur. J. Immunol. 39, 956–964 (2009).
Viret, C. et al. Thymus-specific serine protease contributes to the diversification of the functional endogenous CD4 T cell receptor repertoire. J. Exp. Med. 208, 3–11 (2011).
Nedjic, J., Aichinger, M., Emmerich, J., Mizushima, N. & Klein, L. Autophagy in thymic epithelium shapes the T-cell repertoire and is essential for tolerance. Nature 455, 396–400 (2008).
Nedjic, J., Aichinger, M., Mizushima, N. & Klein, L. Macroautophagy, endogenous MHC II loading and T cell selection: the benefits of breaking the rules. Curr. Opin. Immunol. 21, 92–97 (2009).
Postoak, J. L. et al. Thymic epithelial cells require lipid kinase Vps34 for CD4 but not CD8 T cell selection. J. Exp. Med. 219, e20212554 (2022).
Rodrigues, P. M. et al. LAMP2 regulates autophagy in the thymic epithelium and thymic stroma-dependent CD4 T cell development. Autophagy 19, 426–439 (2023).
Nitta, T. et al. Thymoproteasome shapes immunocompetent repertoire of CD8+ T cells. Immunity 32, 29–40 (2010).
Petrozziello, E. et al. Cathepsin L-dependent positive selection shapes clonal composition and functional fitness of CD4+ T cells. Nat. Immunol. 26, 1127–1138 (2025). This study uses a combination of repertoire sequencing and TCR re-expression in transgenic mice to show that cathepsin L deficiency not only creates gaps in the CD4⁺ T cell repertoire but also alters the functionality of clones whose selection appears cathepsin L independent.
Marrack, P. & Kappler, J. The T cell receptor. Science 238, 1073–1079 (1987).
Tomaru, U. et al. Restricted expression of the thymoproteasome is required for thymic selection and peripheral homeostasis of CD8+ T cells. Cell Rep. 26, 639–651.e632 (2019).
Kincaid, E. Z., Murata, S., Tanaka, K. & Rock, K. L. Specialized proteasome subunits have an essential role in the thymic selection of CD8+ T cells. Nat. Immunol. 17, 938–945 (2016).
Apavaloaei, A. et al. PSMB11 orchestrates the development of CD4 and CD8 thymocytes via regulation of gene expression in cortical thymic epithelial cells. J. Immunol. 202, 966–978 (2019).
Sasaki, K. et al. Thymoproteasomes produce unique peptide motifs for positive selection of CD8+ T cells. Nat. Commun. 6, 7484 (2015).
Hsieh, C. S., deRoos, P., Honey, K., Beers, C. & Rudensky, A. Y. A role for cathepsin L and cathepsin S in peptide generation for MHC class II presentation. J. Immunol. 168, 2618–2625 (2002).
Ohigashi, I., Matsuda-Lennikov, M. & Takahama, Y. Peptides for T cell selection in the thymus. Peptides 146, 170671 (2021).
Ohigashi, I. et al. The thymoproteasome hardwires the TCR repertoire of CD8+ T cells in the cortex independent of negative selection. J. Exp. Med. 218, e20201904 (2021).
Starr, T. K., Jameson, S. C. & Hogquist, K. A. Positive and negative selection of T cells. Annu. Rev. Immunol. 21, 139–176 (2003).
Goncalves, C. M., Henriques, S. N., Santos, R. F. & Carmo, A. M. CD6, a rheostat-type signalosome that tunes T cell activation. Front. Immunol. 9, 2994 (2018).
Azzam, H. S. et al. CD5 expression is developmentally regulated by T cell receptor (TCR) signals and TCR avidity. J. Exp. Med. 188, 2301–2311 (1998).
Matson, C. A. et al. CD5 dynamically calibrates basal NF-κB signaling in T cells during thymic development and peripheral activation. Proc. Natl Acad. Sci. USA 117, 14342–14353 (2020).
Miller, M. L. et al. Basal NF-kappaB controls IL-7 responsiveness of quiescent naive T cells. Proc. Natl Acad. Sci. USA 111, 7397–7402 (2014).
Sinclair, C., Saini, M., Schim van der Loeff, I., Sakaguchi, S. & Seddon, B. The long-term survival potential of mature T lymphocytes is programmed during development in the thymus. Sci. Signal. 4, ra77 (2011).
Xing, Y., Wang, X., Jameson, S. C. & Hogquist, K. A. Late stages of T cell maturation in the thymus involve NF-kappaB and tonic type I interferon signaling. Nat. Immunol. 17, 565–573 (2016).
Takahama, Y. The thymoproteasome in shaping the CD8+ T-cell repertoire. Curr. Opin. Immunol. 83, 102336 (2023).
Xing, Y., Jameson, S. C. & Hogquist, K. A. Thymoproteasome subunit-β5T generates peptide-MHC complexes specialized for positive selection. Proc. Natl Acad. Sci. USA 110, 6979–6984 (2013).
Takada, K. et al. TCR affinity for thymoproteasome-dependent positively selecting peptides conditions antigen responsiveness in CD8+ T cells. Nat. Immunol. 16, 1069–1076 (2015). This work shows that positive selection of monoclonal CD8+ T cells in the absence of the thymoproteasome results in aberrant TCR responsiveness, homeostatic maintenance and immune responses to infection.
Singer, A., Adoro, S. & Park, J. H. Lineage fate and intense debate: myths, models and mechanisms of CD4- versus CD8-lineage choice. Nat. Rev. Immunol. 8, 788–801 (2008).
Hogquist, K. A. & Jameson, S. C. The self-obsession of T cells: how TCR signaling thresholds affect fate ‘decisions’ and effector function. Nat. Immunol. 15, 815–823 (2014).
Vrisekoop, N., Monteiro, J. P., Mandl, J. N. & Germain, R. N. Revisiting thymic positive selection and the mature T cell repertoire for antigen. Immunity 41, 181–190 (2014).
Lucas, B. & Germain, R. N. T-cell repertoire: political correctness in the immune system. Curr. Biol. 6, 783–787 (1996).
Khanom, U. S. et al. TCR affinity for in vivo peptide-induced thymic positive selection fine-tunes TCR responsiveness of peripheral CD8+ T cells. J. Immunol. 203, 881–887 (2019).
Ebert, P. J., Jiang, S., Xie, J., Li, Q. J. & Davis, M. M. An endogenous positively selecting peptide enhances mature T cell responses and becomes an autoantigen in the absence of microRNA miR-181a. Nat. Immunol. 10, 1162–1169 (2009).
Lo, W. L. et al. An endogenous peptide positively selects and augments the activation and survival of peripheral CD4+ T cells. Nat. Immunol. 10, 1155–1161 (2009).
Hogquist, K. A. et al. Identification of a naturally occurring ligand for thymic positive selection. Immunity 6, 389–399 (1997).
Daniels, M. A. et al. Thymic selection threshold defined by compartmentalization of Ras/MAPK signalling. Nature 444, 724–729 (2006). This study uses a series of altered peptide ligands to reveal that a broad spectrum of TCR–self-pMHC affinities supports positive selection.
Hogquist, K. A. et al. T cell receptor antagonist peptides induce positive selection. Cell 76, 17–27 (1994).
Santori, F. R. et al. Rare, structurally homologous self-peptides promote thymocyte positive selection. Immunity 17, 131–142 (2002).
Ashton-Rickardt, P. G., Van Kaer, L., Schumacher, T. N., Ploegh, H. L. & Tonegawa, S. Peptide contributes to the specificity of positive selection of CD8+ T cells in the thymus. Cell 73, 1041–1049 (1993).
Ross, J. O. et al. Distinct phases in the positive selection of CD8+ T cells distinguished by intrathymic migration and T-cell receptor signaling patterns. Proc. Natl Acad. Sci. USA 111, E2550–E2558 (2014).
Shinzawa, M. et al. Unraveling CD8 lineage decisions reveals that functionally distinct CD8+ T cells are selected by different MHC-I thymic peptides. Nat. Immunol. 27, 786–798 (2026).
Stepanek, O. et al. Coreceptor scanning by the T cell receptor provides a mechanism for T cell tolerance. Cell 159, 333–345 (2014).
Clement, M. et al. CD8 coreceptor-mediated focusing can reorder the agonist hierarchy of peptide ligands recognized via the T cell receptor. Proc. Natl Acad. Sci. USA 118, e2019639118 (2021).
Au-Yeung, B. B. et al. Quantitative and temporal requirements revealed for Zap70 catalytic activity during T cell development. Nat. Immunol. 15, 687–694 (2014).
Rogers, D. et al. Pre-existing chromatin accessibility and gene expression differences among naive CD4+ T cells influence effector potential. Cell Rep. 37, 110064 (2021).
Nielsen, H. V. et al. Nr4a1 and Nr4a3 redundantly control clonal deletion and contribute to an anergy-like transcriptome in auto-reactive thymocytes to impose tolerance in mice. Nat. Commun. 16, 784 (2025).
Baptista, A. P. et al. Lymph node stromal cells constrain immunity via MHC class II self-antigen presentation. eLife 3, e04433 (2014).
Roche, P. A. & Furuta, K. The ins and outs of MHC class II-mediated antigen processing and presentation. Nat. Rev. Immunol. 15, 203–216 (2015).
McCarthy, M. K. & Weinberg, J. B. The immunoproteasome and viral infection: a complex regulator of inflammation. Front. Microbiol. 6, 21 (2015).
Mandl, J. N. et al. Quantification of lymph node transit times reveals differences in antigen surveillance strategies of naive CD4+ and CD8+ T cells. Proc. Natl Acad. Sci. USA 109, 18036–18041 (2012).
Seddon, B. & Zamoyska, R. TCR signals mediated by Src family kinases are essential for the survival of naive T cells. J. Immunol. 169, 2997–3005 (2002).
Polic, B., Kunkel, D., Scheffold, A. & Rajewsky, K. How αβ T cells deal with induced TCRα ablation. Proc. Natl Acad. Sci. USA 98, 8744–8749 (2001).
Martin, B., Becourt, C., Bienvenu, B. & Lucas, B. Self-recognition is crucial for maintaining the peripheral CD4+ T-cell pool in a nonlymphopenic environment. Blood 108, 270–277 (2006).
Takada, K. & Jameson, S. C. Self-class I MHC molecules support survival of naive CD8 T cells, but depress their functional sensitivity through regulation of CD8 expression levels. J. Exp. Med. 206, 2253–2269 (2009).
Labrecque, N. et al. How much TCR does a T cell need? Immunity 15, 71–82 (2001).
Surh, C. D. & Sprent, J. Homeostasis of naive and memory T cells. Immunity 29, 848–862 (2008).
Hataye, J., Moon, J. J., Khoruts, A., Reilly, C. & Jenkins, M. K. Naive and memory CD4+ T cell survival controlled by clonal abundance. Science 312, 114–116 (2006).
Vrisekoop, N., Artusa, P., Monteiro, J. P. & Mandl, J. N. Weakly self-reactive T-cell clones can homeostatically expand when present at low numbers. Eur. J. Immunol. 47, 68–73 (2017).
Cho, J. H., Kim, H. O., Surh, C. D. & Sprent, J. T cell receptor-dependent regulation of lipid rafts controls naive CD8+ T cell homeostasis. Immunity 32, 214–226 (2010).
Stefanova, I., Dorfman, J. R. & Germain, R. N. Self-recognition promotes the foreign antigen sensitivity of naive T lymphocytes. Nature 420, 429–434 (2002). This study shows that T cells deprived of access to tonic self-pMHC signals rapidly lose their responsiveness to foreign antigen stimulation.
Spencer, C. T. et al. Viral infection causes a shift in the self peptide repertoire presented by human MHC class I molecules. Proteomics Clin. Appl. 9, 1035–1052 (2015).
Gutierrez-Hoffmann, M. G. et al. Borrelia burgdorferi-induced changes in the class II self-immunopeptidome displayed on HLA-DR molecules expressed by dendritic cells. Front. Med. (Lausanne) 7, 568 (2020).
Mueller, S. N. et al. Regulation of homeostatic chemokine expression and cell trafficking during immune responses. Science 317, 670–674 (2007).
Park, J. et al. Attenuation of migration properties of CD4+ T cells from aged mice correlates with decrease in chemokine receptor expression, response to retinoic acid, and RALDH expression compared to young mice. Biosci. Biotechnol. Biochem. 78, 976–980 (2014).
Kwok, T. et al. Age-associated changes to lymph node fibroblastic reticular cells. Front. Aging 3, 838943 (2022).
Nikolich-Zugich, J., Li, G., Uhrlaub, J. L., Renkema, K. R. & Smithey, M. J. Age-related changes in CD8 T cell homeostasis and immunity to infection. Semin. Immunol. 24, 356–364 (2012).
Thompson, H. L., Smithey, M. J., Surh, C. D. & Nikolich-Zugich, J. Functional and homeostatic impact of age-related changes in lymph node stroma. Front. Immunol. 8, 706 (2017).
Becklund, B. R. et al. The aged lymphoid tissue environment fails to support naive T cell homeostasis. Sci. Rep. 6, 30842 (2016).
Guichard, V. et al. Calcium-mediated shaping of naive CD4 T-cell phenotype and function. eLife 6, e27215 (2017).
Zinzow-Kramer, W. M. et al. Strong basal/tonic TCR signals are associated with negative regulation of naive CD4+ T cells. Immunohorizons 6, 671–683 (2022).
Khan, O. et al. TOX transcriptionally and epigenetically programs CD8+ T cell exhaustion. Nature 571, 211–218 (2019).
Kalekar, L. A. et al. CD4+ T cell anergy prevents autoimmunity and generates regulatory T cell precursors. Nat. Immunol. 17, 304–314 (2016).
Eggert, J. et al. Cbl-b mitigates the responsiveness of naive CD8+ T cells that experience extensive tonic T cell receptor signaling. Sci. Signal. 17, eadh0439 (2024).
Sood, A. et al. Differential interferon-γ production potential among naive CD4+ T cells exists prior to antigen encounter. Immunol. Cell Biol. 97, 931–940 (2019).
Berkley, A. M. & Fink, P. J. Cutting edge: CD8+ recent thymic emigrants exhibit increased responses to low-affinity ligands and improved access to peripheral sites of inflammation. J. Immunol. 193, 3262–3266 (2014).
Moran, A. E. et al. T cell receptor signal strength in Treg and iNKT cell development demonstrated by a novel fluorescent reporter mouse. J. Exp. Med. 208, 1279–1289 (2011).
Smith, K. et al. Sensory adaptation in naive peripheral CD4 T cells. J. Exp. Med. 194, 1253–1261 (2001).
Zinzow-Kramer, W. M., Weiss, A. & Au-Yeung, B. B. Adaptation by naive CD4+ T cells to self-antigen-dependent TCR signaling induces functional heterogeneity and tolerance. Proc. Natl Acad. Sci. USA 116, 15160–15169 (2019).
Hudson, D., Fernandes, R. A., Basham, M., Ogg, G. & Koohy, H. Can we predict T cell specificity with digital biology and machine learning? Nat. Rev. Immunol. 23, 511–521 (2023).
Banerjee, A. et al. T cell receptor cross-reactivity prediction improved by a comprehensive mutational scan database. Cell Syst. 16, 101345 (2025). This study reports a comprehensive cross-reactivity database to predict the peptides that activate a given TCR.
Birnbaum, M. E. et al. Deconstructing the peptide-MHC specificity of T cell recognition. Cell 157, 1073–1087 (2014). Using a large-scale peptide screen, the authors find that many, but not all, peptides recognized by the same TCR share sequence motifs.
Wooldridge, L. et al. A single autoimmune T cell receptor recognizes more than a million different peptides. J. Biol. Chem. 287, 1168–1177 (2012). Using experimental and mathematical analysis, this paper quantified the surprising extent of CD8+ T cell cross-reactivity.
Ekeruche-Makinde, J. et al. Peptide length determines the outcome of TCR/peptide-MHCI engagement. Blood 121, 1112–1123 (2013).
Wucherpfennig, K. W. et al. Polyspecificity of T cell and B cell receptor recognition. Semin. Immunol. 19, 216–224 (2007).
Bhardwaj, V., Kumar, V., Geysen, H. M. & Sercarz, E. E. Degenerate recognition of a dissimilar antigenic peptide by myelin basic protein-reactive T cells. Implications for thymic education and autoimmunity. J. Immunol. 151, 5000–5010 (1993).
Drost, F. et al. Predicting T cell receptor functionality against mutant epitopes. Cell Genom. 4, 100634 (2024).
Kim, Y., Sidney, J., Pinilla, C., Sette, A. & Peters, B. Derivation of an amino acid similarity matrix for peptide: MHC binding and its application as a Bayesian prior. BMC Bioinform. 10, 394 (2009).
Korpela, D. et al. EPIC-TRACE: predicting TCR binding to unseen epitopes using attention and contextualized embeddings. Bioinformatics 39, btad743 (2023).
Moris, P. et al. Current challenges for unseen-epitope TCR interaction prediction and a new perspective derived from image classification. Brief. Bioinform. 22, bbaa318 (2021).
Croce, G. et al. Deep learning predictions of TCR-epitope interactions reveal epitope-specific chains in dual alpha T cells. Nat. Commun. 15, 3211 (2024).
Riley, T. P. et al. T cell receptor cross-reactivity expanded by dramatic peptide-MHC adaptability. Nat. Chem. Biol. 14, 934–942 (2018).
Sewell, A. K. Why must T cells be cross-reactive? Nat. Rev. Immunol. 12, 669–677 (2012).
Vizcaíno, J. A. et al. The human immunopeptidome project: a roadmap to predict and treat immune diseases. Mol. Cell. Proteom. 19, 31–49 (2020).
Robinson, J., Waller, M. J., Parham, P., Bodmer, J. G. & Marsh, S. G. IMGT/HLA database—a sequence database for the human major histocompatibility complex. Nucleic Acids Res. 29, 210–213 (2001).
Granados, D. P., Laumont, C. M., Thibault, P. & Perreault, C. The nature of self for T cells-a systems-level perspective. Curr. Opin. Immunol. 34, 1–8 (2015).
Aro, M. & Joshi R. M. (eds) Handbook of Orthography and Literacy 531–550 (Routledge, 2005).
Cavnar, W. B. & Trenkle, J. M. N-gram-based text categorization. In Proc. SDAIR-94, 3rd Annual Symposium on Document Analysis and Information Retrieval 161–175 (University of Nevada, 1994).
Tiffeau-Mayer, A. et al. How different are self and nonself. PRX Life 4, 013027 (2026).
Achar, S. R. et al. Universal antigen encoding of T cell activation from high-dimensional cytokine dynamics. Science 376, 880–884 (2022).
Oreper, D., Klaeger, S., Jhunjhunwala, S. & Delamarre, L. The peptide woods are lovely, dark and deep: hunting for novel cancer antigens. Semin. Immunol. 67, 101758 (2023).
Phulphagar, K. M. et al. Sensitive, high-throughput HLA-I and HLA-II immunopeptidomics using parallel accumulation-serial fragmentation mass spectrometry. Mol. Cell. Proteom. 22, 100563 (2023).
Gomez-Zepeda, D. et al. Thunder-DDA-PASEF enables high-coverage immunopeptidomics and is boosted by MS2Rescore with MS2PIP timsTOF fragmentation prediction model. Nat. Commun. 15, 2288 (2024).
Zhou, J., Zhang, Y. Y., Li, Q. Y. & Cai, Z. H. Evolutionary history of cathepsin L (L-like) family genes in vertebrates. Int. J. Biol. Sci. 11, 1016–1025 (2015).
Kasahara, M. & Flajnik, M. F. Origin and evolution of the specialized forms of proteasomes involved in antigen presentation. Immunogenetics 71, 251–261 (2019).
van Oers, N. S., Killeen, N. & Weiss, A. ZAP-70 is constitutively associated with tyrosine-phosphorylated TCR zeta in murine thymocytes and lymph node T cells. Immunity 1, 675–685 (1994).
Madrenas, J. et al. Zeta phosphorylation without ZAP-70 activation induced by TCR antagonists or partial agonists. Science 267, 515–518 (1995).
Marangoni, F. et al. The transcription factor NFAT exhibits signal memory during serial T cell interactions with antigen-presenting cells. Immunity 38, 237–249 (2013).
Mingueneau, M. et al. Loss of the LAT adaptor converts antigen-responsive T cells into pathogenic effectors that function independently of the T cell receptor. Immunity 31, 197–208 (2009).
Myers, D. R., Norlin, E., Vercoulen, Y. & Roose, J. P. Active tonic mTORC1 signals shape baseline translation in naive T cells. Cell Rep. 27, 1858–1874.e1856 (2019).
Sood, A. et al. CD5 levels define functionally heterogeneous populations of naive human CD4+ T cells. Eur. J. Immunol. 51, 1365–1376 (2021).
De Simone, G. et al. CXCR3 identifies human naive CD8+ T cells with enhanced effector differentiation potential. J. Immunol. 203, 3179–3189 (2019).
Ashouri, J. F. & Weiss, A. Endogenous Nur77 is a specific indicator of antigen receptor signaling in human T and B cells. J. Immunol. 198, 657–668 (2017).
Burgueno-Bucio, E., Mier-Aguilar, C. A. & Soldevila, G. The multiple faces of CD5. J. Leukoc. Biol. 105, 891–904 (2019).
Bending, D. & Zikherman, J. Nr4a nuclear receptors: markers and modulators of antigen receptor signaling. Curr. Opin. Immunol. 81, 102285 (2023).
Zikherman, J., Parameswaran, R. & Weiss, A. Endogenous antigen tunes the responsiveness of naive B cells but not T cells. Nature 489, 160–164 (2012).
Martin, B. et al. Highly self-reactive naive CD4 T cells are prone to differentiate into regulatory T cells. Nat. Commun. 4, 2209 (2013).
Jergovic, M. et al. Infection-induced type I interferons critically modulate the homeostasis and function of CD8+ naive T cells. Nat. Commun. 12, 5303 (2021).
Ju, Y. J. et al. Self-reactivity controls functional diversity of naive CD8+ T cells by co-opting tonic type I interferon. Nat. Commun. 12, 6059 (2021).
Textor, J. et al. Machine learning analysis of the T cell receptor repertoire identifies sequence features of self-reactivity. Cell Syst. 14, 1059–1073.e5 (2023). Textor et al. (2023) and Lagatutta et al. (2022) performed analyses across large TCR-seq datasets to show that specific TCR sequence features predict cell fate.
Stadinski, B. D. et al. Hydrophobic CDR3 residues promote the development of self-reactive T cells. Nat. Immunol. 17, 946–955 (2016).
Lagattuta, K. A. et al. Repertoire analyses reveal T cell antigen receptor sequence features that influence T cell fate. Nat. Immunol. 23, 446–457 (2022).
Daley, S. R. et al. Cysteine and hydrophobic residues in CDR3 serve as distinct T-cell self-reactivity indices. J. Allergy Clin. Immunol. 144, 333–336 (2019).
Wirasinha, R. C. et al. αβ T-cell receptors with a central CDR3 cysteine are enriched in CD8αα intraepithelial lymphocytes and their thymic precursors. Immunol. Cell Biol. 96, 553–561 (2018).
Jamaleddine, H. et al. Chronic infection control relies on T cells with lower foreign antigen binding strength generated by N-nucleotide diversity. PLoS Biol. 22, e3002465 (2024).
Gavin, M. A. & Bevan, M. J. Increased peptide promiscuity provides a rationale for the lack of N regions in the neonatal T cell repertoire. Immunity 3, 793–800 (1995).
Mora, T. & Walczak, A. M. Towards a quantitative theory of tolerance. Trends Immunol. 44, 512–518 (2023).
Weber, K. S. et al. Distinct CD4+ helper T cells involved in primary and secondary responses to infection. Proc. Natl Acad. Sci. USA 109, 9511–9516 (2012).
ElTanbouly, M. A. et al. VISTA is a checkpoint regulator for naive T cell quiescence and peripheral tolerance. Science 367, eaay0524 (2020).
Deep, D. et al. Precursor central memory versus effector cell fate and naive CD4+ T cell heterogeneity. J. Exp. Med. 221, e20231193 (2024).
Sajani, A. et al. Heterogeneity and plasticity of the naive CD4+ T cell compartment. Cell Rep. 45, 116980 (2026).
Even, Z. et al. The amalgam of naive CD4+ T cell transcriptional states is reconfigured by helminth infection to dampen the amplitude of the immune response. Immunity 57, 1893–1907.e1896 (2024).
Henderson, J. G., Opejin, A., Jones, A., Gross, C. & Hawiger, D. CD5 instructs extrathymic regulatory T cell development in response to self and tolerizing antigens. Immunity 42, 471–483 (2015).
DiToro, D. et al. Differential IL-2 expression defines developmental fates of follicular versus nonfollicular helper T cells. Science 361, eaao2933 (2018).
Lee, V. et al. The endogenous repertoire harbors self-reactive CD4+ T cell clones that adopt a follicular helper T cell-like phenotype at steady state. Nat. Immunol. 24, 487–500 (2023).
Bartleson, J. M. et al. Strength of tonic T cell receptor signaling instructs T follicular helper cell-fate decisions. Nat. Immunol. 21, 1384–1396 (2020).
Lee, G. W. et al. Developmental self-reactivity determines pathogenic Tc17 differentiation potential of naive CD8+ T cells in murine models of inflammation. Nat. Commun. 15, 2919 (2024).
Kim, Y. J. et al. CD5 expression dynamically changes during the differentiation of human CD8+ T cells predicting clinical response to immunotherapy. Immune Netw. 23, e35 (2023).












Leave a Reply