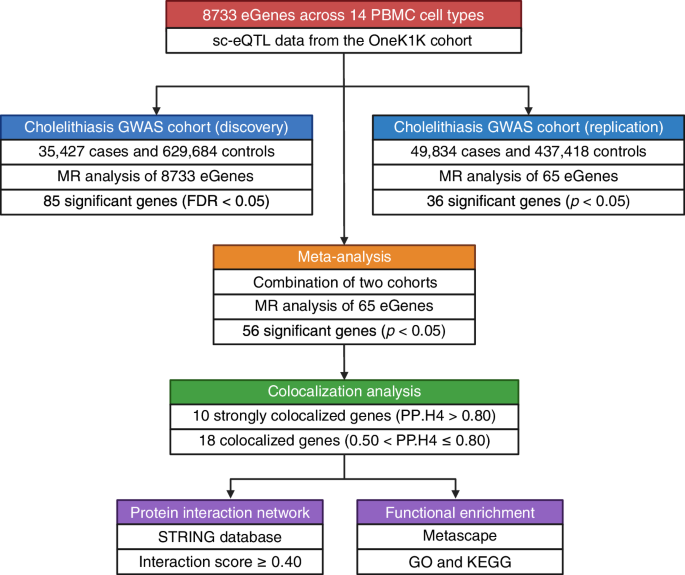
Cholelithiasis, or gallstone disease, is a globally prevalent hepatobiliary disorder imposing a growing burden on healthcare systems, particularly in aging and industrialized societies2. While often clinically silent, symptomatic cases can lead to serious complications such as acute cholecystitis, biliary obstruction, and pancreatitis, requiring invasive interventions21. Despite its high prevalence, the therapeutic landscape for cholelithiasis remains narrow, with surgery as the mainstay treatment and pharmacological options, such as ursodeoxycholic acid, showing limited efficacy and scope7,22. The lack of targeted medical therapies stems in part from insufficient understanding of the disease’s molecular and immunological underpinnings12. In this context, our study addresses a critical gap by leveraging single-cell transcriptomic data, MR, and colocalization analyses to identify immune-cell-specific causal genes associated with cholelithiasis susceptibility. Among the 56 robust eGenes identified, 28 demonstrated both replication and colocalization evidence. Based on the strength of colocalization, 10 eGenes were classified as Tier 1 and 18 eGenes as Tier 2. These eGenes represented the most robust shared genetic associations between immune-cell gene expression and cholelithiasis risk, as they exhibited both replication and strong colocalization evidence. The high-confidence eGenes were further enriched in pathways related to lipid metabolism, immune regulation, and protein ubiquitination, highlighting potential mechanistic links between immune-cell gene regulation and gallstone pathogenesis. These findings highlight key immune regulatory nodes that may represent promising therapeutic entry points for precision therapies aimed at modulating immune cell functions within the gallbladder microenvironment. The following sections delve into how these eGenes, within distinct immune subsets, may orchestrate cellular and molecular pathways that contribute to gallstone pathogenesis.
A particularly striking set of associations emerged within CD4+ T cells, including both CD4 ET and CD4 NC subsets, suggesting that transcriptional regulation in these cells may significantly influence cholelithiasis susceptibility. TMEM258, identified in CD4 ET, is known for its role in endoplasmic reticulum (ER) stress responses and protein glycosylation23. Dysregulation of ER function in CD4 ET may exacerbate pro-inflammatory signaling in the gallbladder, contributing to chronic inflammation and altered biliary secretion24. DAGLB, expressed in CD4 NC, encodes a lipase critical for 2-AG biosynthesis, implicating endocannabinoid signaling in immune-lipid interactions25. Its upregulation could influence T cell activation thresholds and lipid metabolism at the biliary interface. UBE2Q2 and GLYCTK, also elevated in CD4 NC, highlight the potential roles of protein ubiquitination and glycine metabolism in T cell functionality and immune-mediated tissue remodeling26. Interestingly, RGS12 showed divergent effects depending on cell context, positively associated in CD4 ET but negatively in CD4 NC, suggesting its role in modulating T cell signaling may vary by activation state27. Protective associations were observed for UBE2L3, AP1S1, and C2orf74, indicating potential regulatory functions that limit pathological T cell activation or migration28,29. Collectively, these findings point to CD4+ T cells as dynamic mediators of both risk and resilience in gallstone disease, with cell-state-specific gene expression dictating divergent immunological outcomes. The duality of pro- and anti-inflammatory signals within the CD4+ compartment highlights the complexity of immune contributions to cholelithiasis and underscores the need for nuanced, cell-type-targeted therapeutic approaches.
CD8+ T cells, traditionally recognized for their cytotoxic functions, also appear to play an important immunomodulatory role in cholelithiasis. C2orf74, identified in CD8 ET cells, remains poorly characterized in terms of its precise function. Its consistent appearance across both CD4 and CD8 subsets may suggest a fundamental regulatory role in T cell activation or survival. C5orf56 is another gene with unclear function but has been linked to immune cell development, and its upregulation may reflect increased cytotoxic activity or altered T cell homeostasis in the gallbladder microenvironment. PPM1M, a protein phosphatase, may act as a negative regulator of T cell receptor signaling30. Paradoxically, its elevated expression in CD8 ET might either attenuate or dysregulate immune responses depending on context. ITCH, an E3 ubiquitin ligase, has well-established roles in T cell anergy and immune tolerance, but aberrant expression may impair these regulatory mechanisms and exacerbate inflammatory damage to biliary tissues31. Meanwhile, several eGenes showed negative associations, hinting at protective roles. UBE2L3, identified in both CD8 ET and CD8 NC subsets, participates in ubiquitin-mediated protein degradation and immune regulation28. Its reduced expression may disrupt protein turnover or facilitate pathological T cell activation. RPL9, a ribosomal protein, and PHACTR4, a regulator of actin dynamics, may contribute to T cell stability and suppress excessive proliferation or migration. RGS12, again observed in CD8 NC, underscores the importance of G-protein signaling modulation in T cell responses27. Collectively, these findings suggest that CD8+ T cells contribute to gallstone pathogenesis through finely balanced signaling pathways involving cytoskeletal integrity, protein homeostasis, and immune activation thresholds. Disruptions to these regulatory systems may shift CD8+ T cells from surveillance to pathogenic roles, particularly in an inflammatory or cholesterol-rich biliary environment.
In addition to T cells, several non-T immune cell populations, including NK, DC, and B IN, were found to harbor eGenes significantly associated with cholelithiasis, suggesting that the pathogenesis of gallstone disease extends across both adaptive and innate immunity. Among these, TMEM258 also appeared in B IN with a consistently strong association across immune cell subsets, suggesting a potentially shared regulatory role in ER stress and protein folding pathways among immune lineages23. The involvement of RAC1 in NK R points toward cytoskeletal reorganization and migration, critical functions for tissue infiltration and immune surveillance at the biliary interface32. The elevated expression of ITCH and PPM1M in NK may reflect dysregulated ubiquitination and phosphatase activity, potentially altering NK cell-mediated cytotoxicity and cytokine secretion30,33. Interestingly, several eGenes of NK demonstrated protective associations. UBE2L3 and RPL9 may stabilize NK function or limit excessive activation, while DYNLRB1, a dynein light chain component, might contribute to intracellular trafficking and immune synapse formation34. TMEM171, also negatively associated, has unclear function but may play a regulatory role in NK homeostasis. In DC, KCNK17, a two-pore domain potassium channel, suggests ion flux and membrane potential regulation may influence antigen presentation or DC maturation35. Collectively, these findings underscore the involvement of diverse immune effectors in cholelithiasis, with NK in particular balancing pro-inflammatory signaling with regulatory restraint. The discovery of distinct gene signatures across innate immune subsets expands the mechanistic framework of gallstone disease and points toward complex immunometabolic interactions that warrant further functional investigation.
This study offers several key advantages over previous research efforts in the field of cholelithiasis. Foremost among them is the integration of sc-eQTL data, which enables high-resolution mapping of gene expression to specific immune cell subtypes, an advance over bulk eQTL approaches that average signals across heterogeneous tissues36. This cell-type specificity allowed us to identify previously overlooked regulatory genes and pathways that may play causal roles in gallstone disease. Additionally, the integration of MR with Bayesian colocalization provides statistical evidence supporting shared genetic signals between gene expression and gallstone risk, suggesting potential causal relationships that warrant further functional validation16,17. The use of large, well-powered GWAS datasets for both discovery and replication further improves the reliability and generalizability of our findings. Finally, our focus on immune cell subsets, rather than hepatic or metabolic tissues alone, offers a novel immunological perspective on cholelithiasis pathogenesis and uncovers therapeutic targets that may otherwise remain hidden. Together, these innovations establish a comprehensive framework for investigating the immune-genetic architecture of gallstone disease and demonstrate the utility of integrating cutting-edge genomic tools for target discovery.
Despite its strengths, this study has several limitations that should be acknowledged. First, our analysis was restricted to immune-cell-specific eQTLs derived from peripheral blood mononuclear cells (PBMCs) in the OneK1K dataset. As such, non-immune cell types such as hepatocytes, cholangiocytes, and gallbladder epithelial cells were not directly assessed. Although several prioritized eGenes are also broadly expressed in hepatic and metabolic tissues, the current framework cannot capture their potential non-immune regulatory effects. Future studies integrating single-cell or spatial eQTL data from liver and biliary tissues will be essential to elucidate complementary non-immune mechanisms underlying cholelithiasis. Second, while sc-eQTL datasets provide high-resolution insights into cell-type-specific regulation, they remain limited in population diversity, disease representation, and sample size, which may affect the generalizability of our findings across different ancestries and clinical contexts. Additionally, as most sc-eQTL-derived exposures are instrumented by a single SNP, the application of multi-SNP MR models and formal sensitivity analyses was inherently limited. This sparsity reflects the current nature of sc-eQTL resources, where cell-type-specific resolution often comes at the cost of reduced instrument diversity. Third, our replication strategy leveraged updated FinnGen releases. However, partial sample overlap between FinnGen R3 and R12 may reduce the strict statistical independence between discovery and replication stages, and thus replication findings should be interpreted as validation in an expanded cohort rather than a fully non-overlapping replication37. Finally, gene expression does not always correlate with protein abundance or activity, and our study did not directly assess downstream functional consequences at the proteomic or phenotypic levels. Further functional validation will be essential to confirm the causal relationships between the prioritized genes and cholelithiasis.
In summary, this study provides a comprehensive single-cell genetic framework for dissecting the immune contributions to gallstone pathogenesis. Moving forward, future research should integrate multi-ancestry GWAS datasets, spatial transcriptomics, proteomic profiling, and functional assays in relevant hepatobiliary tissues to comprehensively characterize the immunogenetic architecture of cholelithiasis. These efforts will be essential to elucidate the interplay between immune and metabolic pathways and to translate genetic findings into precision diagnostic and therapeutic strategies.













Leave a Reply