Borisov, T., Metodiev, D., Ferdinandov, D. & Roumenina, L. T. Complement system in tumor growth and metastases. Br. J. Cancer https://doi.org/10.1038/s41416-025-03260-6 (2025).
Merle, N. S. & Roumenina, L. T. The complement system as a target in cancer immunotherapy. Eur. J. Immunol. 54, e2350820 (2024).
Rezola Artero, M., Minery, A., Nedelcev, L., Radanova, M. & Roumenina, L. T. Complement and the hallmarks of cancer. Semin. Immunol. https://doi.org/10.1016/j.smim.2025.101950 (2025).
Roumenina, L. T., Daugan, M. V., Petitprez, F., Sautès-Fridman, C. & Fridman, W. H. Context-dependent roles of complement in cancer. Nat. Rev. Cancer 19, 698–715 (2019).
Ajona, D., Cragg, M. S. & Pio, R. The complement system in clinical oncology: Applications, limitations and challenges. Semin. Immunol. 77, 101921 (2025).
Ovcinnikovs, V., Dijkman, K., Zom, G. G., Beurskens, F. J. & Trouw, L. A. Enhancing complement activation by therapeutic anti-tumor antibodies: Mechanisms, strategies, and engineering approaches. Semin. Immunol. 77, 101922 (2025).
Saxena, R., Gottlin, E. B., Campa, M. J., He, Y. W. & Patz, E. F. Complement regulators as novel targets for anti-cancer therapy: a comprehensive review. Semin. Immunol. 77, 101931 (2025).
King, B. C. & Blom, A. M. Intracellular complement: evidence, definitions, controversies, and solutions. Immunol. Rev. 313, 104–119 (2023).
West, E. E. & Kemper, C. Complosome — the intracellular complement system. Nat. Rev. Nephrol. 19, 426–439 (2023).
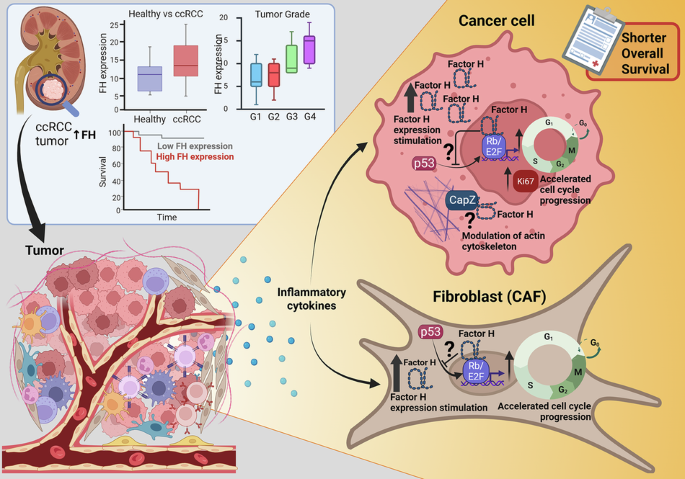
Daugan, M. V. et al. Intracellular factor H drives tumor progression independently of the complement cascade. Cancer Immunol. Res. 9, 909–925 (2021).
Davidson, G. et al. Mesenchymal-like tumor cells and myofibroblastic cancer-associated fibroblasts are associated with progression and immunotherapy response of clear cell renal cell carcinoma. Cancer Res. 83, 2952–2969 (2023).
Saxena, R. et al. Promotion of an antitumor immune program by a tumor-specific, complement-activating antibody. J. Immunol. 212, 1589–1601 (2024).
Saxena, R. et al. Complement factor H: a novel innate immune checkpoint in cancer immunotherapy. Front. Cell Dev. Biol. 12, 1–9 (2024).
Clarke, J. M. et al. Complement factor H targeting antibody GT103 in refractory non-small cell lung cancer: a phase 1b dose escalation trial. Nat. Commun. 16, 93 (2025).
Bond, K. H. et al. The extracellular matrix environment of clear cell renal cell carcinoma determines cancer associated fibroblast growth. Cancers 13, 1–27 (2021).
Yap, N. Y. et al. Establishment of epithelial and fibroblast cell cultures and cell lines from primary renal cancer nephrectomies. Cell Biol. Int. 43, 715–725 (2019).
Karlsson, M. et al. A single–cell type transcriptomics map of human tissues. Sci. Adv. 7, 1–9 (2021).
Lin, H. et al. Mutual activation between cancer-associated fibroblasts and cancer cells facilitates growth and progression of gastric cancer. Int. J. Clin. Exp. Pathol. 13, 2691–2700 (2020).
Liu, T. et al. Conditional knockout of telomerase reverse transcriptase in mesenchymal cells impairs mouse pulmonary fibrosis. PLoS ONE 10, 1–16 (2015).
Kosugi, S., Hasebe, M., Tomita, M. & Yanagawa, H. Systematic identification of cell cycle-dependent yeast nucleocytoplasmic shuttling proteins by prediction of composite motifs. Proc. Natl. Acad. Sci. USA 106, 10171–10176 (2009).
Hellwage, J. et al. Complement C3b/C3d and cell surface polyanions are recognized by overlapping binding sites on the most carboxyl-terminal domain of complement factor H. J. Immunol. 169, 6935–6944 (2002).
Leffler, J. et al. Annexin-II, DNA, and histones serve as factor H ligands on the surface of apoptotic cells. J. Biol. Chem. 285, 3766–3776 (2010).
Jokiranta, T. S., Hellwage, J., Koistinen, V., Zipfel, P. F. & Meri, S. Each of the three binding sites on complement factor H interacts with a distinct site on C3b. J. Biol. Chem. 275, 27657–27662 (2000).
Smyrek, I. et al. E-cadherin, actin, microtubules and FAK dominate different spheroid formation phases and important elements of tissue integrity. Biol. Open 8 https://doi.org/10.1242/bio.037051 (2019).
Hu, J. et al. Multi-omic profiling of clear cell renal cell carcinoma identifies metabolic reprogramming associated with disease progression. Nat. Genet. 56, 442–457 (2024).
Becht, E. et al. Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression. Genome Biol. 17, 1–20 (2016).
Merle, N. S., Church, S. E., Fremeaux-Bacchi, V. & Roumenina, L. T. Complement system part I – Molecular mechanisms of activation and regulation. Front. Immunol. 6, 262 (2015).
Revel, M. et al. Humoral complementomics–exploration of noninvasive complement biomarkers as predictors of renal cancer progression. Oncoimmunology 13 https://doi.org/10.1080/2162402X.2024.2328433 (2024).
Roumenina, L. T. et al. Tumor cells hijack macrophage-produced complement C1q to promote tumor growth. Cancer Immunol. Res. 7, 1091–1105 (2019).
De Vivar Chevez, A. R., Finke, J. & Bukowski, R. The role of inflammation in kidney cancer. Adv. Exp. Med. Biol. 816, 197–234 (2014).
Kamińska, K., Czarnecka, A. M., Escudier, B., Lian, F. & Szczylik, C. Interleukin-6 as an emerging regulator of renal cell cancer. Urol. Oncol. Semin Orig. Investig. 33, 476–485 (2015).
Chen, Q. et al. Growth-induced stress enhances epithelial-mesenchymal transition induced by IL-6 in clear cell renal cell carcinoma via the Akt/GSK-3β/β-catenin signaling pathway. Oncogenesis 6 https://doi.org/10.1038/oncsis.2017.74 (2017)
Nguyen, T. N., Nguyen-Tran, H. H., Chen, C. Y. & Hsu, T. IL6 and CCL18 mediate cross-talk between VHL-deficient kidney cells and macrophages during development of renal cell carcinoma. Cancer Res. 82, 2716–2733 (2022).
Elvington, M., Liszewski, M. K., Bertram, P., Kulkarni, H. S. & Atkinson, J. P. A C3(H20) recycling pathway is a component of the intracellular complement system. J. Clin. Invest. 127, 970–981 (2017).
Ault, B. H. et al. Human factor H deficiency: mutations in framework cysteine residues and block in H protein secretion and intracellular catabolism. J. Biol. Chem. 272, 25168–25175 (1997).
Fenaille, F. et al. Site-specific N-glycan characterization of human complement factor H. Glycobiology 17, 932–944 (2007).
Merle, D. A. et al. mTOR inhibition via rapamycin treatment partially reverts the deficit in energy metabolism caused by FH loss in RPE cells. Antioxidants 10, 1–18 (2021).
Crosby, M. E. & Almasan, A. Opposing roles of E2Fs in cell proliferation and death. Cancer Biol. Ther. 3, 1208–1211 (2004).
Fogal, V., Hsieh, J. K., Royer, C., Zhong, S. & Lu, X. Cell cycle-dependent nuclear retention of p53 by E2F1 requires phosphorylation of p53 at Ser315. EMBO J. 24, 2768–2782 (2005).
Hsieh, J. K. et al. Novel function of the cyclin A binding site of E2F in regulating p53-induced apoptosis in response to DNA damage. Mol. Cell Biol. 22, 78–93 (2002).
Gardner, W. D. & Hoch, S. O. Binding specificity of the two major DNA-binding proteins in human serum. J. Biol. Chem. 254, 5238–5242 (1979).
Gardner, W. D., White, P. J. & Hoch, S. O. Identification of a major human serum DNA-binding protein as β1H of the alternative pathway of complement activation. Biochem. Biophys. Res. Commun. 94, 61–67 (1980).
Edwards, M. et al. Capping protein regulators fine-tune actin assembly dynamics. Nat. Rev. Mol. Cell Biol. 15, 677–689 (2014).
Park, S. et al. Emerging roles of TRIO and F-actin-binding protein in human diseases. Cell Commun. Signal 16, 2–6 (2018).
Seldeen, K. L. et al. Absence of complement factor H reduces physical performance in C57BL6 mice. Immunobiology 225, 152003 (2020).
Mahajan, S. et al. Local complement factor H protects kidney endothelial cell structure and function. Kidney Int. 100, 824–836 (2021).
Alexander, J. J. et al. Absence of complement factor H alters bone architecture and dynamics. Immunobiology 223, 761–771 (2018).
Meri, S. Self-nonself discrimination by the complement system. FEBS Lett. 590, 2418–2434 (2016).
Boudhabhay, I. & Roumenina, L. T. Complement factor H: a guardian within?. Kidney Int. 100, 747–749 (2021).
Armento, A. et al. Loss of complement factor H impairs antioxidant capacity and energy metabolism of human RPE cells. Sci. Rep. 10, 1–15 (2020).
Li, Y. et al. Glomerular mesangial cells derived complement factor H regulates complement activation, influences cell proliferation, and maintains actin cytoskeleton. Int. Immunopharmacol. 154, 114544 (2025).
Amornsiripanitch, N. et al. Complement factor H autoantibodies are associated with early stage NSCLC. Clin. Cancer Res. 16, 3226–3231 (2010).
Bushey, R. T. et al. A therapeutic antibody for cancer, derived from single human B cells. Cell Rep. 15, 1505–1513 (2016).
Bushey, R. T. et al. Antitumor immune mechanisms of the anti-complement factor H antibody GT103. Mol. Cancer Ther. 22, 778–789 (2023).
Campa, M. J., Gottlin, E. B., Bushey, R. T. & Patz, E. F. Complement Factor H antibodies from lung cancer patients induce complement-dependent lysis of tumor cells, suggesting a novel immunotherapeutic strategy. Cancer Immunol. Res. 3, 1325–1332 (2015).
Clarke, J. M. et al. Complement factor H targeting antibody GT103 in refractory non-small cell lung cancer: a phase 1b dose escalation trial. Nat Commun. 16, 93 (2025)
Hu, J. et al. dataset from Multi-omic profiling of clear cell renal cell carcinoma identifies metabolic reprogramming associated with disease progression. Nat. Genet. 56, 442–457 (2024).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. ClusterProfiler: An R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Liberzon, A. et al. The molecular signatures database hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Hu, C. et al. CellMarker 2.0: an updated database of manually curated cell markers in human/mouse and web tools based on scRNA-seq data. Nucleic Acids Res. 51, D870–D876 (2023).
Sun, D. et al. TISCH: A comprehensive web resource enabling interactive single-cell transcriptome visualization of tumor microenvironment. Nucleic Acids Res. 49, D1420–D1430 (2021).
Sun, D. et al. Datasets from TISCH: A comprehensive web resource enabling interactive single-cell transcriptome visualization of tumor microenvironment: UCEC_GSE139555, STAD_GSE134520, SKCM_GSE115978, PAAD_CRA001160, OV_GSE118828, NSCLC_GSE131907, NHL_GSE128531, NET_GSE140312, MCC_GSE117988, LIHC_GSE125449, KIRC_GSE111360, HNSC_GSE103322, CRC_GSE146771, CHOL_GSE125449, BRCA_GSE138536 and BCC_GSE123813. Nucleic Acids Res. 49, D1420–D1430 (2021). http://tisch.comp-genomics.org/.
Ramos, M. et al. Multiomic integration of public oncology databases in bioconductor. JCO Clin. Cancer Informatics. 4, 958–971 (2020).
Meylan, M. et al. Dataset from: Tertiary lymphoid structures generate and propagate anti-tumor antibody-producing plasma cells in renal cell cancer: GSE175540. Immunity 55, 527–541.e525 (2022).
Dragon-Durey, M. A. et al. Heterozygous and homozygous factor H deficiencies associated with hemolytic uremic syndrome or membranoproliferative glomerulonephritis: report and genetic analysis of 16 cases. J. Am. Soc. Nephrol. 15, 787–795 (2004).
Wu, J. et al. Structure of complement fragment C3b-factor H and implications for host protection by complement regulators. Nat. Immunol. 10, 728–733 (2009).
Boldt, K., van Reeuwijk, J., Gloeckner, C. J., Ueffing, M. & Roepman, R. Tandem affinity purification of ciliopathy-associated protein complexes. Methods Cell Biol. 91, 143–160 (2009).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Szklarczyk, D. et al. The STRING database in 2023: protein-protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 51, D638–D646 (2023).
Morgan, H. P. et al. Structural basis for engagement by complement factor H of C3b on a self surface. Nat. Struct. Mol. Biol. 18, 463–471 (2011).
Schmidt, C. Q. et al. A new map of glycosaminoglycan and C3b binding sites on factor H. J. Immunol. 181, 2610–2619 (2011).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 1–21 (2014).















Leave a Reply