Mice
Specific pathogen-free male C57BL/6 mice (8–10 weeks old) were purchased from Beijing Vital River Laboratory Animal Technology. Mice were maintained in a specific pathogen-free animal facility (temperature: 21–25 °C; humidity: 30–70%; 12-h light/12-h dark cycle) at the State Key Laboratory of Biotherapy, Sichuan University. All animal experiments were approved by the Ethics Review Committee of Animal Care and Use Committee of Sichuan University (ethics number: 20230307045).
Human lung tissue
The studies were conducted in accordance with the principles of the Declaration of Helsinki and were approved by the Ethics Committee of West China Hospital, Sichuan University (approval number: 2016-98). Written informed consent was obtained from all participants before inclusion in the study. No participant compensation was provided.
Human lung tissues were obtained from two sources. Fibrotic lung tissues were obtained at the time of lung transplantation from individuals with a diagnosis of IPF (n = 8, all men, median age 66.5, range 51–70). Non-IPF lung tissues (n = 4, all men, median age 64.5, range 50–67) were obtained from uninvolved regions from individuals undergoing lobectomy for lung and esophageal carcinomas. All tissues were fresh-frozen immediately after surgical resection and stored at −80 °C before processing. Detailed demographic information (sex, age and clinical diagnosis) for all donors is provided in Supplementary Table 1.
The human lung tissue samples used in this study were fully depleted or destroyed during the immunopeptidome profiling experiments (including tissue homogenization, lysis, immunoprecipitation, peptide elution and MS analysis). No remaining tissue samples or derived material (for example, peptides, lysates or MHC complexes) were conserved for further analysis.
Participant recruitment
Participants were not actively recruited for this study. Instead, fibrotic lung tissues were opportunistically obtained from consecutive individuals undergoing lung transplantation for end-stage IPF at West China Hospital, Sichuan University, between 2023 and 2025, where tissue collection was part of routine clinical biobanking with prior ethics approval and written informed consent. Non-IPF control lung tissues were obtained from uninvolved (distal paracancerous) regions of lobectomy specimens from individuals undergoing surgery for lung or esophageal carcinoma at the same institution during the same period. Tissue collection occurred only when surplus material was available after routine pathological assessment and only after written informed consent had been obtained for research use.
All IPF cases met the American Thoracic Society and European Respiratory Society diagnostic criteria for definite or probable IPF, as determined by multidisciplinary discussion. No additional inclusion/exclusion criteria beyond clinical indication for transplantation or lobectomy were applied for this study.
Immunopeptidome data processing of sample preparation and MS analysis
Control lung and lung fibrosis tissues were first homogenized in a grinding mill with liquid nitrogen. Subsequently, lysis buffer was added to the pellet and incubated on ice for 1 h. After incubation, the mixtures were sonicated for 3 min on ice and centrifuged at 20,000g at 4 °C for 1 h. The human HLA class I peptide complexes were enriched using a NeoDiscovery immunopeptidome enrichment kit (BZNEO-H-I-10, BaizhenBio), which targets HLA-A/HLA-B/HLA-C. Similarly, mouse MHC class I peptide complexes were enriched using a NeoDiscovery kit (BZNEO-M-I-10, BaizhenBio), which targets H-2 class I. The supernatant was carefully aspirated for the enrichment step of the HLA class I–peptide complex with an antibody to HLA class I provided in the kit. The immunopeptides were further separated and desalted using a C18 monospin column from GL Science. The eluted fractions containing the immunopeptides were then dried using a speed vacuum instrument. The peptides were reconstituted in 0.1% formic acid and separated on a homemade C18 column (200 mm × 75 μm inner diameter, 1.9-Å particle size; Dr. Maisch) using a nano-Elute HPLC system (Bruker) at a flow rate of 300 nl min−1 with a 60-min gradient of 4–100% acetonitrile (containing 0.1% formic acid). The data were acquired using a timsTOF Pro2 mass spectrometer (Bruker) in data-dependent acquisition mode with 10 parallel accumulation–serial fragmentation events. The mass range was set from 300 to 1,700 m/z, and the ion threshold for MS2 was set to 2,500. The acquired MS/MS data were analyzed using PEAKS Studio (v11.5) against reviewed protein databases downloaded from UniProt (www.uniprot.org), including the human database (20,201 entries) and the canonical mouse database (17,167 entries). For the identification of the immunopeptidome, we used a built-in peptidome workflow with the following parameters: the tolerance for precursor and fragment mass were set to 20 ppm and 0.05 Da, respectively. Oxidation of methionine (+15.9949 Da) was set as a variable modification. The enzyme was set as none. A contaminant database containing 147 proteins was added for excluding the contaminant peptides. False discovery rate of the identified peptides was set at 1%. Confident amino acid threshold was set as 2%.
MHC class I binding predictions
MHC class I binding predictions were determined using NetMHCpan35 (v.4.0).
Positional frequency matrices of peptides
Motifs identified by GibbsCluster36 (v.2.0) were summarized as position frequency matrices P of size 9 × 20, whose rows sum to 1. Positional frequency matrices of MHC class I-presented peptides were transformed into 180-dimensional vectors to calculate pairwise correlations, which were then used for hierarchical clustering based on Euclidean distance, principal component analysis for dimensionality reduction and visualization and linear correlation analysis with HLA class I allele overlap, enabling assessment of motif separation across samples.
The information content at each motif position Pi (i = 1 ~ 9) was quantified using Shannon entropy (Hi), representing the number of bits required to define the amino acid distribution (j = A, C, D, …, Y) at that position:
$${H}_{i}\,=\,-{\varSigma }_{{j}}\,{P}_{ij}\,{\log }_{2}({P}_{ij}).$$
scRNA-seq data processing
Chromium single-cell data were aligned to the GRCh38 genome reference using cellranger (v.7.1.0). The CellRanger count pipeline was used to generate feature–barcode matrices, which were essential to downstream clustering and gene expression analysis. Cell quality was assessed by two standards: (1) cells with expressed genes over 4,000 or less than 600 were filtered, and (2) cells with more than 10% mitochondrial counts among CD45+ cells were filtered out, as were those with more than 70% mitochondrial counts among CD45− cells. In addition, cells with multiple cell markers were identified as doublets. After quality control, the BLM-induced mouse dataset contained 20,433 single cells, the human dataset contained 243,472 single cells, and the peptide-treated (MAF116–124, APBB270–78, TNS3119–127) mouse dataset contained 77,169 single cells, all of which were available for downstream analysis.
Dimensionality reduction and clustering
After quality control of cells and genes, dimensionality reduction and unsupervised clustering were performed by using the Seurat37 (v.4.0.1) R package. First, we normalized the unique molecular identifier (UMI) count matrix using the SCTransform function (regressing out the effects of percentages of mitochondria). The RunPCA function was used to calculated principal components on the scaled data. The top 30 principal components were defined as the input of the next step according to the Elbow plot. A graph-based clustering approach was applied to cluster these cells. We used the FindNeighbors function to construct a shared nearest neighbor graph and refine edge weights between two cells. We then applied the Louvain algorithm to iteratively group cells together with the FindClusters function. For visualization, the dimensionality of each dataset was further reduced using UMAP with the Seurat function RunUMAP. When calculating the UMAP coordinates, the effective scale of embedded points setting varied from 0.2 to 1.2.
Functional enrichment analysis
Functional enrichment of Gene Ontology terms was assessed using the clusterProfiler R package. Differentially expressed genes with an adjusted P value of <0.05 were used as input.
Comparison of macrophage and fibroblast cell subsets with human datasets
To compare the macrophages and fibroblasts in our BLM-induced mouse dataset to those previously published for individuals with fibrosis and healthy individuals (GEO GSE136831), we extracted the UMI matrices for all relevant cells. After reclustering high-quality cells from the human dataset, we manually annotated cell types based on characteristic genes (Extended Data Fig. 5). To systematically compare pathological cells across different human individuals and mouse models, we projected cells between datasets based on the expression of homologous genes characteristic of humans and mice (Fig. 4c).
Define phenotype characterization of cell subsets
Enrichment scores for immunopeptide sets corresponding to each cell type were calculated using the score.cells.puram function from pagoda2, with the source gene set as input. To define macrophage characterization of M2 phenotypes, we collected gene signatures for M2 (anti-inflammatory/alternatively activated) phenotype macrophages38, including Ccl4, Ccl13, Ccl18, Ccl22, Cd276, Ctsa, Ctsb, Ctsc, Ctsd, Fn1, Il4r, Irf4, Lyve1, Mmp9, Mmp14, Mmp19, Msr1, Tgfb1, Tgfb2, Tgfb3, Tnfsf8, Tnfsf12, Vegfa, Vegfb and Vegfc. We then applied a gene set enrichment-based supervised classification method using the AddModuleScore function to calculate the phenotype score for each macrophage.
Establishment of the Fib-SCORE pipeline for potential antigen with therapeutic effect prioritization
To identify peptides with antifibrotic therapeutic potential, there were two major challenges that needed addressing: the high interindividual variability of human HLA class I recognition and the complexity of MHC class I antigen presentation across distinct cell populations. To overcome these obstacles while simplifying the experimental system, we developed Fib-SCORE, a scoring-based pipeline using the BLM-induced mouse model as a proof-of-concept platform. Fib-SCORE integrates stringent quality control, presentation stability, allelic binding potential and targeting specificity to enable the systematic discovery of therapeutic peptides. This framework was designed to conceptually validate the hypothesis that MHC class I ligands can serve as actionable therapeutic targets in fibrosis, while preserving the key pathological features of fibrotic antigen presentation.
The pipeline begins with a dedicated filtering step to generate the candidate peptide repertoire. Each peptide is required to meet the following four criteria: (1) peptide identification confidence (evaluated using MS/MS spectrum matching quality, ΔRT, MS2 correlation and related metrics), (2) predicted binding affinity (with an affinity (nM) of <1,000), (3) source protein conservation (requiring that the peptides are derived from source proteins with mapped homologs in humans) and (4) sample presentation stability (requiring detection in more than half of samples and expression of the corresponding source gene in matched scRNA-seq datasets). These filters not only reduce noise but also facilitate the prioritization of fibrosis-associated targets with potential relevance to human fibrosis, thereby enhancing the biological interpretability of our findings.
Building on this filtered set, Fib-SCORE implements a ranking strategy that defines therapeutic peptide antigens according to the following three criteria: (1) specificity score (peptide expression abundance differential; prioritizes peptides that are both disease specific and stably presented within groups, reducing peptides restricted to individual HLA alleles), (2) affinity score (predicted peptide–MHC class I binding affinity; prioritizes peptides with high allelic binding potential) and (3) target score (potential targeting of pathological cell populations; prioritizes peptides that target homologous pathological cell types in both humans and mice). Each score is calculated using the following equations.
First, based on differential peptide expression, the specificity score of each peptide is calculated using equation (1)
$$\mathrm{Specificity}\,\mathrm{score}=\left\{\begin{array}{cc}\left(1-\,P\right)* \mathrm{sgn}({\log }_{2}\mathrm{FC}), & \mathrm{shared}\\ 1, & \mathrm{BLM-specific}\end{array}\right)$$
(1)
where P and log2FC represent the P value and log2 (fold change) of peptide differential expression (BLM versus PBS) significance calculated using the limma R package, respectively.
Second, the affinity score is computed according to
$$\mathrm{Affinity}\,\mathrm{score}\,=\,1-\frac{{ \% }\rm{Rank}}{100}$$
(2)
where %Rank is the prediction values transformed to percentile rank scores of NetMHCpan. The final peptide predicted affinity corresponds to the one with the highest score of a favorable HLA allele.
Third, the target score of each peptide based on scRNA-seq data (source gene) is calculated using equation (3)
$$\mathrm{Target}\,\mathrm{score}\,=\,\left\{\begin{array}{cc}1, & \mathrm{no}\,\mathrm{expression}\\ {\mathrm{TS}}_{\mathrm{{Candidate}|{Enrichment}}\,}, & \mathrm{other}\end{array}\right)$$
(3)
where TSCandidate|Enrichment is defined as 1 − (P value), where the P value is computed using Fisher’s exact test (2 × 2 contingency table, rows of the table distinguish between features categorized as ‘Candidate’ and ‘Noncandidate’, whereas the columns represent ‘Enrichment’ and ‘Unenrichment’), which reflects the specificity level of gene expression in candidate cell types. The following are explanations of the candidate and enrichment categories: (1) candidate indicates the key fibrosis-relevant cell types reported in the pulmonary fibrosis literature25,26,27 and identified in single-cell datasets, including mesothelial cells, endothelial cells, macrophages and fibroblasts, and (2) enrichment indicates the condition of average difference (avg_diff) between a specific cell type and other cell types, as determined by running the FindAllMarkers function on the scaled data. If avg_diff > 0, we infer that the gene is likely to be enriched within that specific cell group compared to other cell types.
Finally, the score S for each peptide was calculated by combining the specificity score, affinity score and target score as follows:
$$S=\mathrm{Specificity}\,\mathrm{score}\,+\,\mathrm{Affinity}\,\mathrm{score}\,+\,\mathrm{Target}\,\mathrm{score}.$$
Mouse experimental procedures
To establish the BLM-induced pulmonary fibrosis model, C57BL/6 mice were anesthetized with 5% isoflurane and then intratracheally instilled with 3 mg per kg (body weight) BLM (MCE) in 50 μl of PBS per mouse using a 29-gauge insulin syringe.
To assess the therapeutic effects of peptide vaccine in the BLM-induced pulmonary fibrosis model, peptide vaccine (50 μg per mouse) was intramuscularly injected into C57BL/6 mice on days 0, 7 and 14. Mice were killed 28 days after the instillation of BLM for the isolation and processing of major organs and serum samples.
Lung index analysis
Lung index analysis represents a straightforward method for estimating the extent of pulmonary edema. A detailed description of the methodology is provided as follows. On the final day of the experiment, intact lung tissue was excised and weighed. The lung index was then calculated using the following formula:
Lung index = lung weight (g) / body weight (g)
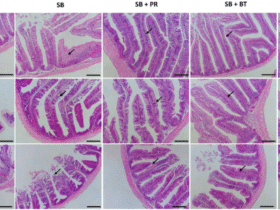
From the representative morphology of the fibrotic tissue (below), the extent of pulmonary fibrosis was correlated with lung weight and body weight. In the current study, lung index serves as a critical parameter in evaluating vaccine efficacy.
Preparation of peptide vaccine
The preparation process of liposome-based peptide vaccines involves three main steps: preparation of the 1V209 and hydrogenated soybean phosphatidylcholine mixture stock solution, microfluidic encapsulation and ultrafiltration39.
In the first step, the TLR7 stock solution was prepared. The total amount of peptide required was calculated to be 1.5 mg for the entire process. To achieve this, 1.5 mg of 1V209 was weighed and placed in a 1.5-ml Eppendorf tube, where it was dissolved in 80 μl of DMSO. This mixture was subjected to a water bath and sonication until it became completely clear. Concurrently, 7–8 mg of hydrogenated soybean phosphatidylcholine (egg lecithin) was weighed and dissolved in 150 μl of anhydrous ethanol. To this solution, 36 μl of cholesterol solution, at a concentration of 30 μg μl−1, was added. The resulting mixture was then transferred to the Eppendorf tube containing 1V209, and anhydrous ethanol was added to bring the total volume to 300 μl, yielding a final concentration of 5 μg μl−1 for 1V209.
In the second step (microfluidic encapsulation), the water bath was preheated to 60 °C to prepare the system. The organic phase was prepared by diluting 60 μl of the stock solution with anhydrous ethanol to a final volume of 375 μl, achieved by adding 315 μl of ethanol. Simultaneously, the aqueous phase was prepared by adding 75 μl of a 10 mg ml−1 peptide solution to water, bringing the final volume to 1,125 μl. These two phases were then mixed using a micro and nano microfluidic system at 37 °C, ensuring thorough encapsulation.
The final step involved ultrafiltration, where the microfluidic mixture was subjected to ultrafiltration at 4,000 rpm for 20 min. This process effectively removed water, organic solvent and any unencapsulated peptides, resulting in the completed preparation of the liposome-based peptide vaccines.
TCR sequencing
Extraction of total RNAs from the samples was accomplished using TRIzol reagent (Invitrogen, 15596018). This process adhered to the extraction protocols as described by Chomczynski et al.40. Subsequent to RNA extraction, a DNA digestion process was executed using DNaseI (Thermo Scientific, EN0521).
To ascertain RNA quality, the ratio of absorbance at 260 nm to 280 nm was evaluated using a Nanodrop OneC spectrophotometer (Thermo Scientific, ND-ONEC-W). RNA integrity was further verified through 1.5% agarose gel electrophoresis. RNA samples that met the quality criteria were subsequently quantified using a Qubit 3 fluorometer (Invitrogen, Q33216), along with a Qubit RNA HS Assay kit (Invitrogen, Q32855).
For preparation of the TCR sequencing library, approximately 2 μg of total RNA from each sample was used. This was done using a KC-Digital Stranded TCR-seq Library Prep kit for Illumina 150 (Seqhealth Technology, DT0813-02). The procedure was performed in accordance with the manufacturer’s instructions. The kit is designed to minimize duplication bias during PCR and sequencing by incorporating UMIs, which are 8-base random sequences, to tag the preamplified cDNA molecules. The library products, sized between 250 and 500 base pairs, were selected, quantified and ultimately sequenced using a NovaSeq platform from Illumina.
RNA-seq data analysis for TCR sequencing
The initial step in processing the raw sequencing data involved using SOAPnuke (v.1.6.0) to filter out low-quality reads and those that contained adapter sequence contamination. Subsequently, the refined clean reads were subjected to custom script-based processing to address and remove any duplication biases that may have been introduced during the library preparation and sequencing phases. To further refine the data and minimize errors and biases associated with the sequencing process, UMIs were implemented.
The process can be summarized as follows. Clean reads were initially grouped based on their UMI sequences, creating clusters of reads that shared identical UMI sequences. Within these clusters, reads were compared using pairwise alignment, and those with a sequence similarity exceeding 95% were selected to form a new subset of clusters. Once all such subclusters were established, a multiple sequence alignment was conducted for each to derive a single consensus sequence.
These deduplicated consensus sequences served as the foundation for the subsequent TCR-sequencing analysis and were aligned to the globally recognized ImMunoGeneTics database41 using MiXCR software (v.3.0.3). This alignment process was crucial for accurate extraction of the CDR3 sequences, which are pivotal for understanding the adaptive immune response. The R package immunarch (v.0.9.0) was used for statistical analysis and results presentation, including CDR3 length distribution, diversity indices and clonotype tracking.
Flow cytometry analysis
For preparation of lung samples, lungs were minced and digested with 1 mg ml−1 collagenase type I (Gibco) + 0.5 mg ml−1 collagenase type IV (Gibco) in RPMI 1640 at 37 °C and 200 rpm for 45 min, filtered through a 70-μm strainer and washed with PBS, and red blood cell lysis was achieved with red blood cell lysis buffer (154 mM NH4Cl, 10 mM KHCO3, 0.1 mM EDTA 2Na (pH 7.4)). Cells were washed with PBS twice and resuspended in PBS for subsequent analysis.
For cell surface staining, cells (1 × 106 cells per 100 μl per tube) were stained with 1 μl of antibodies at 4 °C for 30 min in the dark. Cells were then washed and resuspended in PBS (500 μl per tube). The following fluorophore-conjugated antibodies purchased from BioLegend were used to evaluate MHC class I expression in immune and stromal cells from lungs of mice 28 days after PBS or BLM treatment: PerCP-Cyanine5.5-conjugated anti-CD45 (103132), PE-conjugated anti-Ly6G (127608), PE-Cyanine7-conjugated anti-Siglec-F (155528), BV421-conjugated anti-CD11c (117343), FITC-conjugated anti-CD64 (139316), BV510-conjugated anti-Ly6C (128033), BV650-conjugated anti-I-A/I-E (107641), APC-conjugated anti-H-2Kᵇ (16518), BV421-conjugated anti-CD326 (118225) and PE-conjugated anti-vimentin (699310). BV510-conjugated anti-CD31 (63089) was purchased from BD Biosciences, and FITC-conjugated anti-α-SMA (F3777) was purchased from Sigma-Aldrich.
To evaluate cytotoxic CD8⁺ T cells in lung tissues of mice 21 days after BLM administration, with or without MAF116–124 vaccine treatment, cells were first stained for surface markers, including PerCP‑Cyanine5.5-conjugated anti‑CD3 (100218), BV421‑conjugated anti‑CD4 (100531) and APC‑conjugated anti‑CD8 (100712), at 4 °C for 30 min in the dark. Subsequently, cells were fixed and permeabilized using a Fixation/Permeabilization kit (554715), followed by intracellular staining with PE/Cyanine7‑conjugated anti‑mouse IFNγ (163508) and PE‑conjugated anti‑mouse/human granzyme B antibodies (372207) for flow cytometric analysis. All fluorophore-conjugated antibodies were purchased from BioLegend.
For the preparation of lymph node samples, lymph nodes were triturated, filtered through a 70-μm strainer and washed with PBS twice. Cells were resuspended in PBS for subsequent analysis. To detect antigen-specific CD8⁺ T cells, single-cell suspensions (1 × 106 cells per tube) were prepared from spleens of immunized and nonimmunized mice. Cells were incubated for 30 min at room temperature with PE-conjugated H-2Db peptide–MHC class I tetramers specific for MAF116–124 (ALISNSHQL) and APBB270–78 (YALTNIQAA) and H-2Kb peptide–MHC class I tetramers specific for TNS3119–127 (SSYMHFTNV), which were custom synthesized by BetterGen. Following tetramer incubation, cells were stained with PerCP-Cy5.5-conjugated anti-CD3, APC-conjugated anti-CD8 and BV421-conjugated anti-CD4. After washing, samples were analyzed on a NovoCyte flow cytometer with NovoExpress 1.4.1 software (ACEA Biosciences).
IHC
To evaluate fibrosis in lung tissues, IHC staining for α-SMA and collagen I was performed. Lung or liver tissues were collected, fixed in 4% paraformaldehyde overnight at 4 °C and embedded in paraffin. Four-micron-thick sections were deparaffinized in xylene and rehydrated through a graded ethanol series. Antigen retrieval was performed by heating sections in sodium citrate buffer (pH 6.0) at 95 °C for 15 min, followed by cooling to room temperature. Endogenous peroxidase activity was blocked with 3% hydrogen peroxide for 10 min.
Tissue sections were then blocked with 5% bovine serum albumin (BSA) for 1 h and incubated overnight at 4 °C with primary antibodies to α-SMA and collagen I (Servicebio). After washing with PBS, sections were incubated with horseradish peroxidase-conjugated secondary antibodies for 1 h at room temperature. Color development was performed using a diaminobenzidine substrate kit (Vector Laboratories), followed by counterstaining with hematoxylin. Sections were dehydrated, mounted with a neutral resin and imaged using Pannoramic MIDI (3DHISTECH). For quantitative analysis, the percentage of α-SMA+ and collagen I+ area was measured using IHC profiler.
H&E staining
H&E staining was performed to assess tissue morphology in lung sections. Paraffin-embedded tissue sections (4 μm thick) were deparaffinized in xylene and rehydrated through a descending ethanol series. Sections were stained with hematoxylin (Vector Laboratories) for 5 min, followed by differentiation in 1% acid alcohol and rinsing in running tap water. Eosin staining was then performed for 2 min, and sections were dehydrated through ascending ethanol concentrations, cleared in xylene and mounted with a neutral resin. Images were captured using Pannoramic MIDI (3DHISTECH), and lung histopathological changes were evaluated using the Szapiel score, which evaluates inflammatory infiltration, alveolar wall thickening and fibrotic lesions.
Masson’s trichrome staining
Masson’s trichrome staining was conducted to assess collagen deposition in lung fibrosis. Tissue sections were deparaffinized and rehydrated, followed by incubation in Bouin’s solution at 56 °C for 1 h for tissue fixation enhancement. After cooling and washing in running tap water, sections were sequentially stained with Weigert’s iron hematoxylin for 10 min, Biebrich scarlet acid fuchsin for 10 min and phosphotungstic/phosphomolybdic acid for 10 min to differentiate collagen fibers. Sections were then stained with aniline blue for 5 min, washed and treated with 1% acetic acid for 2 min. Finally, sections were dehydrated, cleared in xylene and mounted. Collagen fibers appeared blue, whereas muscle and cytoplasm stained red. Lung fibrosis severity was quantified using the Ashcroft score, which evaluates collagen deposition and fibrosis progression.
Sirius Red staining
Sirius Red staining was performed to detect collagen fiber content in lung tissues. Tissue sections were deparaffinized, rehydrated and incubated in Sirius Red solution (0.1% Sirius Red in saturated picric acid) for 1 h at room temperature. Excess dye was removed by washing in 0.5% acetic acid for 2 min. Sections were then dehydrated in absolute ethanol, cleared in xylene and mounted. Collagen fibers appeared red under brightfield microscopy and exhibited birefringence under polarized light.
Immunofluorescence staining
Lung specimens were fixed with 4% formaldehyde and embedded in paraffin. Sections (3 µm) were deparaffinized and then subjected to an EDTA buffer for antigen retrieval. The sections were incubated with blocking buffer (5% goat serum) for 30 min at room temperature and subsequently incubated with primary antibodies overnight at 4 °C, which were prepared in 1% BSA in PBS. Next, the sections are incubated with secondary antibodies at room temperature for 1 h. Finally, the cell nucleus was stained with 1 μg ml−1 DAPI for 5 min at room temperature. Sections were mounted with fluorescent mounting medium (Sigma-Aldrich, F6182) and observed with a Vectra 3 automated quantitative pathology imaging system (Akoya Biosciences). Overlap coefficiency was analyzed using ImageJ.
The primary antibodies used include Cy3-conjugated rabbit anti-mouse MAF (1:50, Bioss), rat anti-mouse F4/80 (1:50, Abcam) and FITC mouse anti-mouse α-SMA (1:500, Sigma). The secondary antibody used was goat anti-mouse IgG-AF488 (1:1,000, Life Science).
In addition, formalin-fixed paraffin-embedded mouse lung tissue sections (4 μm) were subjected to multicolor immunofluorescence staining (cyclic multiplexed tyramide signal amplification) using a IRISKit HyperView mTSA kit (MH900804, Chengdu LUMINIRIS Bio Tech). After antigen retrieval, the tissue sections were sequentially incubated with primary antibodies, including anti-perforin (31647, Cell Signaling Technology, 1:100), anti-cleaved caspase-3 (9579, Cell Signaling Technology, 1:400), anti-α-SMA (IRS094RT, HUABIO, 1:100), anti-CD8 (IRS083, HUABIO, 1:100) and anti-granzyme B (ab255598, Abcam, 1:500). Between each antibody incubation, the sections were treated with IRISKit HyperView Advanced Ab-Stripping buffer (MH900103) to remove bound antibodies, and fluorescence quenching was performed with IRISKit HyperView quench buffer (MH900102) between staining cycles to minimize spectral overlap. After the final staining step, nuclei were counterstained with DAPI. Whole-slide images were acquired using an Evident VS200 slide scanning system. Image alignment and multispectral analysis were performed with IRRISSin software (LUMINIRIS).
Histological scoring of lung inflammation and fibrosis
Pulmonary inflammation was semiquantitatively assessed in H&E-stained sections using the Szapiel scoring system42. This method grades the extent of alveolitis based on the proportion of lung tissue affected, as observed by light microscopy. Specifically, the following scoring criteria were used: grade 0, no alveolitis (normal alveolar structure); grade 1, mild alveolitis with preserved architecture and involvement of <25% of the lung; grade 2, moderate alveolitis involving 25–50% of the lung and grade 3, severe alveolitis involving >50% of the lung. For each sample, scores were determined by examining at least ten nonoverlapping fields at ×200 magnification, with the final score representing the average across these fields.
Pulmonary fibrosis was semiquantitatively evaluated in Masson’s trichrome-stained sections using the Ashcroft scoring scale43. This system assigns grades from 0 to 8 based on the severity and extent of fibrotic changes in the lung parenchyma. Specifically, grade 0 indicates normal lung, grades 1–2 indicate minimal to moderate fibrous thickening of alveolar or bronchiolar walls without significant architectural damage, grades 3–4 indicate moderate thickening with some damage to lung structure and formation of fibrous bands, grades 5–6 indicate severe fibrosis with definite distortion of architecture and large fibrous areas, and grades 7–8 indicate very severe fibrosis with near-total or total fibrous obliteration of the field. Scores were averaged from at least ten random fields per section at ×200 magnification.
To overcome potential bias, all histological samples were coded with unique identifiers and blinded to treatment groups, animal identifiers and experimental conditions before evaluation. Scoring was performed independently by three trained observers (one board-certified pathologist and two research associates not involved in the study design or execution). Interobserver discrepancies (occurring in <10% of cases) were resolved through joint review and consensus. The scorers were distinct from the personnel conducting animal treatments, sample collection and downstream data analyses, ensuring independence. Statistical analysis of the scores was performed by a separate team member after unblinding.
Transmission electron microscopy
Mouse lung tissues were fixed in 2.5% glutaraldehyde, followed by postfixation in 1% osmium tetroxide. The samples were dehydrated through a graded acetone series (30%, 50%, 70%, 80%, 90%, 95% and three changes of 100%) and infiltrated with mixtures of acetone and Epon-812 resin at ratios of 3:1, 1:1 and 1:3 sequentially. Finally, the samples were embedded in pure Epon-812 resin and polymerized. Semithin sections were first examined under a light microscope to localize the regions of interest in mouse lung tissues. Ultrathin sections (60–90 nm) were then prepared using an ultramicrotome and mounted onto copper grids. Sections were stained with uranyl acetate for 10–15 min and lead citrate for 1–2 min at room temperature. Images were acquired using a JEM-1400FLASH transmission electron microscope (JEOL).
ELISpot assay
Lymphocytes were isolated from the spleens of 6- to 8-week-old male C57BL/6 mice using mouse lymphocyte separation medium (Dakewe Biotech), and splenocytes were cocultured with 10 μg ml−1 MAF116–124 for 18 h, after which secretion of IFNγ was measured by ELISpot according to the manufacturer’s instructions (ELD5819, R&D Systems, Diaclone). The number of IFNγ+ spots was counted under a dissection microscope.
Adoptive transfer of T cells
MAF116–124– and PBS-vaccinated mice were killed and their splenic T cells were isolated, as previously described44,45. Briefly, the spleens were digested and passed through a 70-μm strainer. Afterward red blood cells were lysed with ACK lysis buffer (118-156-101, Quality Biology), and the remaining splenocytes were washed with PBS. T cells were then isolated using a pan-T cell isolation kit (STEMCELL Technologies, 19851). In total, 1 × 106 T cells in DMEM were intravenously administered at 3, 6 and 9 days after BLM treatment. Mice were killed 28 days after the instillation of BLM for the isolation and processing of lung tissue.
Molecular docking
The structure model of H-2Db (PDB 4HUU) was obtained from the Protein Data Bank, and the ligand structure of peptide MAF116–124 (‘ALISNSHQL’) was modeled using AlphaFold2. The protein structures were prepared using ChimeraX and PyMOL. The peptide MAF was docked into the structure using the ZDOCK module of Discovery Studio (v.19). In brief, the PDB accession numbers of receptor proteins and ligand were input into the dialog box, and the sampling angle was configured to 6°. The clustering section was expanded, and the parameters for root mean square deviation cutoff, interface cutoff and maximum number of clusters were modified to 6.0, 9.0 and 60, respectively. ZRank was set to a false state, as this parameter does not incorporate factors such as electrostatic potential energy and van der Waals forces. After completing the task, the docking poses in each cluster were examined, and the pose with the highest score was selected for subsequent data analysis. The structures of the homologous proteins, namely HLA-A*02:01, HLA-C*03:04, HLA-B*13:01 and HLA-A*24:02, were predicted using AlphaFold2. The results of the molecular docking were visualized using Discovery Studio.
Induction of peptide-specific CTLs
PBMCs were isolated from fresh buffy coats of healthy HLA-A*02:01+ volunteer donors by density gradient centrifugation using Ficoll-Paque PLUS. All donors provided written informed consent. Adherent monocytes were enriched by plating PBMCs at 2 × 106 cells per ml in RPMI 1640 medium supplemented with 10% heat-inactivated fetal bovine serum (FBS) in six-well plates for 2 h at 37 °C with 5% CO2. Nonadherent cells were gently removed by washing three times with warm PBS and cryopreserved in 90% FBS + 10% DMSO for later use as autologous responder PBLs. Adherent monocyte-enriched cells were differentiated into immature moDCs in RPMI 1640 + 10% FBS containing 10 ng ml−1 recombinant human IL-4 and 1,000 U ml−1 recombinant human GM-CSF at 37 °C with 5% CO2. Half of the medium was replaced with fresh cytokine-containing medium every 2 days. After 6 days, immature moDCs (1 × 106 cells per well per group) were loaded with 20 µg ml−1 MAF116–124 (sequence ALISNSHQL; >95% purity) in 1 ml of RPMI 1640 + 10% FBS for 24 h at 37 °C with 5% CO2.
Peptide-loaded moDCs were then matured by adding a cytokine cocktail consisting of 10 ng ml−1 recombinant human TNF, 10 ng ml−1 recombinant human IL-1β, 1,000 U ml−1 recombinant human IFNγ, 250 ng ml−1 prostaglandin E2, 1 µg ml−1 R848 and 20 µg ml−1 poly(I:C) for 20–24 h at 37 °C with 5% CO2.
Mature peptide-loaded moDCs were collected, washed twice with PBS and cocultured with autologous nonadherent PBLs (responder cells) at a DC:responder ratio of 1:10 (1 × 105 moDCs + 1 × 106 PBLs per well) in 2 ml of RPMI 1640 + 10% FBS in six-well plates. After 24 h, recombinant human IL-2 (5 ng ml−1), IL-7 (5 ng ml−1) and IL-15 (10 ng ml−1) were added to the culture. Half of the medium was replaced with fresh medium containing the same cytokines every 3 days. Seven days after the initial coculture, cultures were restimulated with freshly prepared peptide-loaded mature moDCs at the same 1:10 ratio. This restimulation cycle was repeated weekly for a total of three cycles (three restimulations). After the third cycle, expanded MAF116–124-specific CTLs were collected and used for downstream assays, including IFNγ ELISpot and cytotoxicity assays.
CD8+ T cell proliferation assay
CTLs were labeled with CFSE (Invitrogen) at a final concentration of 1.5 μM for 20 min at 37 °C. After washing, the CFSE-prelabeled CTLs were restimulated by mature DCs (1 × 105 cells per group) loaded with MAF116–124 for 72 h. Next, cells were collected and stained with BV510-labeled anti-CD8 for 30 min for further analysis by flow cytometry. The proliferation percentage of CD8+ T cells was analyzed according to the reduction in CFSE staining intensity.
Cytotoxicity assay
The HLA-A*0201-positive, TAP-1-deficient human hybrid lymphoblast cell line T2 (KL-C1793H, Shanghai Kanglang Biotechnology) and macrophages and myofibroblasts derived from lung tissue of individuals with IPF were used as target cells.
T2 cells were incubated with 3 μg ml−1 β2-microglobulin and MAF116–124 or unrelated peptide (50 μg ml−1) for 16 h at 37 °C with 5% CO2. In vitro cytotoxicity assays were performed with CTLs and peptide-loaded T2 cells. Patient-derived lung M2 macrophages and myofibroblasts are induced from macrophages and fibroblasts isolated from IPF lung tissue with human IL-4 (20 ng ml−1) and TGFβ (10 ng ml−1) for 24 h.
Target cells were washed and labeled with CFSE (Abcam). Following the washing steps, CTLs were cocultured with target cells at effector:target ratios of 1:1, 1:3 and 1:9 for 6 h at 37 °C with 5% CO2 in RPMI 1640 medium supplemented with 10% FBS and 50 U ml−1 penicillin–streptomycin. Each experimental condition was performed in three replicates. Cells were then washed with PBS containing 0.5% BSA (Miltenyi Biotec, 130-091-376) and labeled with 7-AAD (Abcam), according to the manufacturer’s protocol, before data acquisition using a NovoCyte flow cytometer (ACEA Biosciences), and the results were analyzed using NovoExpress software. The target cells were identified as CFSE+, and the dead target cells were identified as CFSE+7-AAD+. Spontaneous death was defined as the proportion of dead target cells cultured alone (negative control), and this value was subtracted from the proportion of dead target cells cocultured with effector cells.
Statistics and reproducibility
For the bioinformatics analysis of human and mouse data, statistical analysis was performed using R software (v.4.3.1). To identify differentially expressed genes between two clusters in the scRNA-seq data, we used wilcox.test in the FindAllmarkers function to evaluate the significance of each gene, with multiple hypothesis correction using the Benjamini–Hochberg procedure. Genes and peptides with an adjusted Pvalue of <0.05 were considered differentially expressed. Peptide differential expression (BLM versus PBS) significance was calculated using the limma R package. The log2 (fold change) for each gene was calculated by subtracting the log2-transformed mean count in each group.
No statistical methods were used to predetermine sample sizes. In the animal studies, mice were randomly assigned to different experimental groups before treatment. Data collection and analysis were not performed blind to the conditions of the experiments. No data were excluded from the analysis except for animals that died during BLM model induction. All mouse experiments were independently repeated at least twice with similar results. Data distribution for both animal and cell-based experiments was assumed to be normal but this was not formally tested. In mouse experiments, data are presented as mean values with standard deviation. Statistical analyses were performed using GraphPad Prism (v.8.0.2) software. Significance was determined by one-way ANOVA for comparisons involving a single independent variable and two-way ANOVA for experiments involving two independent variables, as indicated in each figure legend. Comparisons between two groups were analyzed using two-tailed unpaired Student’s t-tests. P values of <0.05 were considered statistically significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.












Leave a Reply