Ticks and saliva collection
Rhipicephalus microplus ticks (Porto Alegre strain, free from Babesia spp. and Anaplasma spp.) were obtained from a laboratory colony maintained as previously described53. Hereford calves were acquired from a tick-free area and infested with 10-day-old R. microplus larvae. Fully engorged female ticks were obtained after the spontaneous detachment from the calves. Ticks were rinsed with sterile water, dried on a paper towel, and placed dorsal side down on a glass slide containing tape. Salivation was induced by injecting 5 μl of 2% pilocarpine hydrochloride (Sigma-Aldrich, St. Louis, Missouri, USA), dissolved in PBS (pH 7.4), into the ventral side of the lower right coxa using a Hamilton syringe (Hamilton Company, Reno, Nevada, USA). Rm-saliva was collected using a pipette tip from ticks maintained at 28 °C in a humid chamber for approximately 3 h. Rm-saliva was pooled in each collection and the same pooled saliva preparation was used for all experiments in this study. Collected Rm-saliva was stored at −80 °C until use. Animals used in these experiments were housed in Faculdade de Veterinária, Universidade Federal do Rio Grande do Sul (UFRGS), Brazil. Experiments were conducted considering ethical and methodological guidelines in agreement with the Norms by the Animal Experimentation Ethics Committee of the UFRGS. The protocols 27559 and 45209 were approved by the Comissão de Ética no Uso de Animais (CEUA)-UFRGS. We have complied with all relevant ethical regulations for animal use.
Blood collection
Peripheral blood from clinically healthy adult cattle (Holstein breed, female) was collected at the Field Science Center for the Northern Biosphere, Hokkaido University (Sapporo, Hokkaido, Japan). PBMCs were separated from the blood samples using density-gradient centrifugation on Percoll (GE Healthcare, Chicago, Illinois, USA). The experiments using bovine blood samples were approved by the Ethics Committee of the Faculty of Veterinary Medicine, Hokkaido University (approval numbers 17-0024 and 22-0038).
Cell preparation
To isolate CD14+ cells from bovine PBMCs, PBMCs were incubated with anti-bovine CD14 monoclonal antibody (mAb) (CAM36A; Washington State University Monoclonal Antibody Center, Pullman, Washington, USA) for 30 min at 25 °C. Cells were then incubated with anti-mouse IgG1 MicroBeads (Miltenyi Biotec, Bergisch Gladbach, Germany) for 30 min at 25 °C. After incubation, CD14+ cell were sorted using MS columns and MiniMACS Separator (Miltenyi Biotec). CD14⁻ PBMCs were prepared from negative fractions of CD14+ cell sorting. The purity of the CD14+ and CD14⁻ cells (>90%) was confirmed using a spectral cell analyzer SA3800 (SONY, Tokyo, Japan).
PBMC culture
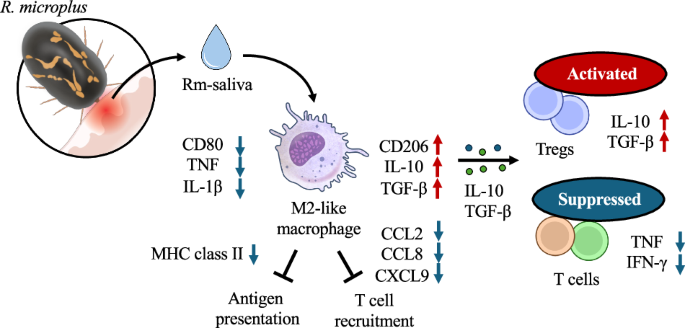
All cells were suspended in RPMI1640 medium (Sigma–Aldrich) supplemented with 10% heat-inactivated FBS (Thermo Fisher Scientific, Waltham, Massachusetts, USA), 200 IU/mL penicillin, 200 μg/mL streptomycin, and 2 mM L-glutamine (Thermo Fisher Scientific). PBMCs and CD14⁻ PBMCs were cultured with 100-fold diluted Rm-saliva for 24 h. After incubation, culture supernatants were collected and the concentration of IL-10 was measured by ELISA, and the cells were subjected to the expression analysis of immune checkpoint molecules or intracellular IL-10 and TGF-β by flow cytometry. In another set of experiments, PBMCs and CD14⁻ PBMCs were first cultured with 100-fold diluted Rm-saliva for 24 h, followed by stimulation with either 1 μg/mL anti-bovine CD3 mAb (MM1A, Washington State University Monoclonal Antibody Center) and 1 μg/mL anti-bovine CD28 mAb (CC220, Bio-Rad, Hercules, California, USA), or 20 μg/ml ConA (Sigma-Aldrich) at 37 °C under 5% CO2 for an additional 24 h. Supernatants were collected, and the concentrations of TNF and IFN-γ were measured by ELISA, and the cells were subjected to the expression analysis of intracellular TNF and IFN-γ.
CD14+ cell culture
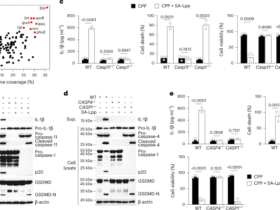
CD14⁺ cells isolated by magnetic sorting were differentiated into macrophages by culturing with bovine M-CSF (50 ng/mL)54 for 6 days, with medium replacement every 48 h. After differentiation, the cells were stimulated with 100-fold diluted Rm-saliva in the presence of LPS (1 μg/mL; Sigma-Aldrich) and bovine IFN-γ (25 ng/mL; Kingfisher Biotech, Saint Paul, Minnesota, USA) for 24 h. Culture supernatants were collected, and the concentrations of TNF, IL-1β, and IL-10 were measured by ELISA. The cells were subjected to the analysis of macrophage phenotyping by flow cytometry or conducted RNA extraction for transcriptomic analysis.
ELISA
To evaluate cytokine production, culture supernatants of PBMCs, CD14⁻ PBMCs or macrophages differentiated from CD14+ cells were collected, and the concentrations of IFN-γ, TNF, IL-10, and IL-1β were measured in duplicate. IFN-γ and TNF were quantified using the Bovine IFN-γ ELISA Development Kit (Mabtech, Nacka Strand, Sweden) and the ELISA Flex: Bovine TNF-α (Mabtech). IL-1β was measured using the Bovine IL-1β ELISA Reagent Kit (Thermo Fisher Scientific), and IL-10 was measured by sandwich ELISA using anti-bovine IL-10 mAb (CC318; Bio-Rad) as the capture antibody and biotinylated anti-bovine IL-10 mAb (CC320; Bio-Rad) as the detection antibody. The concentrations of chemokines in macrophage culture supernatants were determined using the BOVINE CCL2, CCL3, CCL4, CCL5, CCL8, and CXCL9 DO-IT-YOURSELF ELISA Kits (Kingfisher Biotech). Absorbance was measured at 450 nm using a microplate reader (MTP-900; Corona Electric, Tokyo, Japan). For multivariate analysis, raw values of concentration of each cytokine and chemokine were log-transformed, z-score–standardized and subjected to PCA and hierarchical clustering, using R version 4.4.2 with the packages dplyr, ggplot2, and pheatmap.
Flow cytometry
Flow cytometry was used to analyze immune inhibitory cytokine expression in CD4⁺ T cells, immune checkpoint molecule expression in T cells, macrophage polarization, and MHC expression on macrophages. The antibodies used for staining are listed in Table S1, and gating strategies are shown in Supplementary Figs. 1, 6, 8, and 9. To block nonspecific binding to Fc receptors, cells (1 × 10⁶) were preincubated with PBS containing 10% heat-inactivated goat serum (Thermo Fisher Scientific) at 25 °C for 15 min. Cells were then stained with surface antibodies at 25 °C for 20 min. After staining, cells were washed with PBS containing 0.5% bovine serum albumin (Sigma-Aldrich), 2 mM disodium EDTA (Dojindo Molecular Technologies, Kumamoto, Japan), and 0.1% sodium azide (Sigma-Aldrich). For intracellular cytokine staining, Brefeldin A (10 µg/mL) was added 6 h before staining. After surface staining, cells were fixed and permeabilized using BD Cytofix/Cytoperm solution (BD Biosciences, Franklin Lakes, New Jersey, USA) and then stained for intracellular cytokines. For Foxp3 staining in Treg analysis, the Foxp3 Fix/Perm Buffer Set (BioLegend, San Diego, California, USA) was used. The anti-bovine PD-1, TIM-3, LAG-3 and CTLA-4 mAbs used in this study were generated as previously described55,56,57,58. Samples were analyzed using an LSRFortessa flow cytometer (BD Biosciences) or a spectral cell analyzer SA3800 (SONY). Data analysis was conducted using FlowJo v10.4 software (BD Biosciences) or the SA3800 software (SONY).
RNA-seq
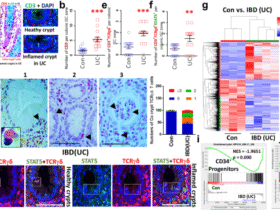
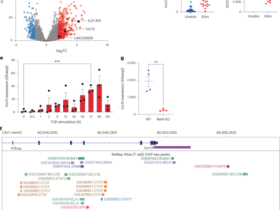
Total RNA was extracted from macrophages using Direct-zol RNA Miniprep Kit (Zymo Research, Irvine, California, USA). Non-stranded RNA-seq libraries were prepared using SMART-Seq v4 Ultra Low Input RNA Kit (Takara Bio, Kusatsu, Japan). Sequencing was carried out with a length of 2 × 150 bp paired-end reads on a NovaSeq X Plus (Illumina, San Diego, California, USA) and generated 2× 13.3 million raw paired-end reads per library. Raw data of RNA-seq were quality-checked by using FastQC version 0.11.9, and trimming was conducted using fastp version 20.1. Mapping was performed with Salmon version 1.10.0 using the ARS-UCD1.3 cDNA index for cattle. In this genome assembly, some bovine immune-related genes (e.g., HLA-DRA) are annotated with projected human gene symbols. For clarity and consistency with bovine immunological nomenclature, gene symbols were curated manually by cross-referencing the ARS-UCD1.2 annotation. Transcript quantification data were imported into R version 4.4.2 using the tximport package, and differential gene expression analysis was conducted with DESeq2. Volcano plots displayed differentially expressed genes between PBS and Rm-saliva affected macrophages, with significance thresholds set at log2 (fold change) > |0.58| and adjusted p-value < 0.05. Heatmaps showed the relative expression (z-score) of the top 80 upregulated and downregulated genes. GO enrichment analysis was performed using the clusterProfiler package to identify significantly enriched biological processes, molecular functions, and cellular components among differentially expressed genes, with adjusted p-value < 0.05 considered significant. GSEA of macrophages was conducted using KEGG pathway gene sets, comparing gene expression data from PBS and Rm-saliva–treated macrophages with the combined data from other fractions. Pathways with false discovery rate (FDR) < 0.25 were defined as enriched. KEGG pathway analysis was performed to illustrate functional enrichment of genes increased or decreased in macrophages, using a significance threshold of adjusted p-value < 0.05.
Immunofluorescence staining of skin sections
Skin samples were collected from two individual adult cattle within the same herd, including tick-infested and non-infested. In tick-infested cattle, a skin specimen was collected from the site with actively feeding ticks. The ticks attached to skin were in the stage of full engorgement (approximately 20 days old). A control sample was obtained from non-infested cattle of similar age and same breed. Tissues were fixed in 10% paraformaldehyde, embedded in paraffin, and sectioned at 5 µm thickness. Sections were deparaffinized, rehydrated through graded ethanol, and subjected to antigen retrieval using heat-induced epitope retrieval in citrate-based buffer. After blocking with 10% goat serum-containing buffer (Nichirei Bioscience; Tokyo Japan) to reduce non-specific binding, tissue sections were incubated with primary antibodies against CD3 (CD3-12, Thermo Fischer Scientific), Iba-1 (Fujifilm Wako, Tokyo Japan), and IL-10 (CC318, Bio-Rad) at 4 °C overnight. Anti-CD3 antibody was conjugated with Alexa Fluor 488 and anti-IL-10 antibody was conjugated by Zenon anti-mouse IgG1 Alexa Fluor 647 (Thermo Fischer Scientific). After washing, Alexa Fluor 555 conjugated goat anti-rabbit IgG (H+L) (Thermo Fischer Scientific) was applied for 1 h at room temperature. Nuclei were counterstained with Hoechst. Stained sections were mounted using VECTASHIELD antifade mounting medium (Vector Laboratories; Newark, California, USA) and observed by confocal microscopy (Zeiss LSM 700; Carl Zeiss, Jena, Germany), and the obtained images were analyzed using Zen software (Carl Zeiss). Ten fields were observed in each experiment.
H&E staining
Skin sections were also subjected to standard H&E staining. Briefly, sections were stained with hematoxylin to visualize nuclei, rinsed, differentiated, and blued, followed by eosin staining to visualize cytoplasmic and extracellular structures. After dehydration and clearing, sections were mounted with a permanent mounting medium. H&E-stained sections were examined using a bright-field microscope, and representative images were acquired from comparable regions of the dermis in tick-nonattached and tick-attached skin.
Statistics and reproducibility
Statistical analysis was performed using R version 4.3.1. For comparisons among four related groups—(i) PBMCs and CD14⁻ PBMCs treated with either PBS or saliva, and (ii) macrophages stimulated with PBS, saliva, with or without LPS and IFN-γ—the Friedman test followed by the Wilcoxon signed-rank test with Holm correction was used. P values less than 0.05 were considered statistically significant. Sample sizes (n) represent independent biological replicates derived from individual animals unless otherwise indicated. Experiments were repeated independently to confirm reproducibility.
Inclusion & ethics statement
This study was conducted through collaborative research involving researchers from multiple institutions in accordance with relevant institutional and national guidelines. All authors contributed to the study and approved the final manuscript.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.














Leave a Reply