Selection of TCRs
The 16 disease-derived TCRs analyzed in this study were previously obtained and characterized from AS and AAU samples using previously reported approaches. For clarity, they fall into three categories, outlined below.
Motif-guided TRBV9+ enrichment (public family)
For TCR 4.1, T cells from the peripheral blood of an individual with HLA-B*27+ AS were expanded in vitro for 28 days using anti-CD3 and IL-2 followed by flow sorting on TRBV9+ CD8 T cells for single-cell TCR sequencing (scTCRseq)24. For TCRs 8.4 and 9.1, TRBV9+ CD8 T cells from the synovial fluid of individuals with HLA-B*27+ AS were flow-sorted and subjected to scTCRseq24. These belong to the well-described BV9-Y/FSTDTQ-BJ2.3/TRAV21 public family.
Non-BV9 example (broadened motif)
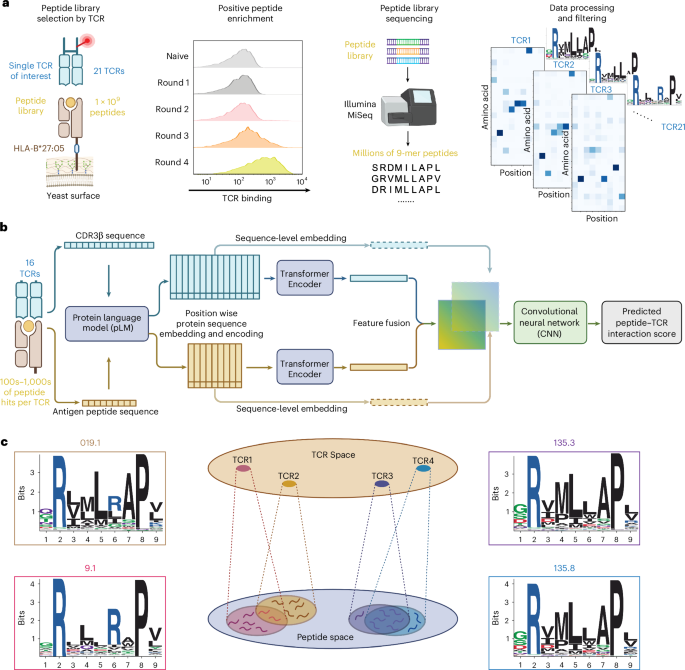
For TCR 019.1, immune cells from the aqueous humor of a participant with HLA-B*27+ AAU were subjected to scTCRseq24. This clonotype does not carry the BV9-Y/FSTDTQ-BJ2.3 motif but instead represents an alternative β-chain background paired with TRAV21, broadening the family beyond BV9 usage.
YeiH tetramer-guided isolation
For TCRs 19.2, 26.1, 26.2, 28.1, 135.1, 135.3, 135.4, 135.7 and 135.8, T cells from the peripheral blood of HLA-B*27+ AAU participants were expanded in vitro for 8 days using YeiH-pulsed autologous peripheral blood mononuclear cells followed by flow sorting on YeiH tetramer+ CD8 T cells for scTCRseq26. These include both TRBV9 and TRBV5 clonotypes (TRBV5-1 and TRBV5-5, IMGT nomenclature), ensuring diversity beyond the canonical BV9 family.
Engineered TCR variants and α-chain swaps
To assess model generalization within a local neighborhood, we generated five engineered variants of TCR 19.2 by introducing one to three amino acid substitutions into its CDR3β loop while maintaining the original α-chain. These engineered receptors (19.2C1–19.2C5) were expressed and profiled in parallel, providing a controlled neighborhood of closely related clonotypes for assessing model generalization.
To validate the β-centric modeling approach, we generated chimeric TCR constructs by swapping AJ28-matched α-chains. The α-chain sequences of TCRs 8.4 and 135.4 were synthesized and cloned into a pHR lentiviral vector containing the WT 19.2 β-chain. These chimeric receptors were expressed in SKW-3 cells (DSMZ, ACC53) and assayed for activation alongside the parental WT TCRs.
Flow cytometry using PSG5–HLA-B27:05 tetramers to stain CD8 T cells from AAU, AS and healthy controls
Study approval
All human participants were enrolled in accordance with Declaration of Helsinki principles and using protocols approved by the institutional review board of Washington University in St. Louis (Institutional Review Board 201912043 and 202212047). Written informed consent was received from participants before inclusion in the study.
Participants
Participants with HLA-B27+ AAU fulfilled the Standardization of Uveitis Nomenclature Working Group classification criteria for HLA-B27+ AAU30. Participants with HLA-B27+ AS satisfied the modified New York criteria31. Two individuals with AAU also met diagnosis of nonradiographic axial spondyloarthritis by Assessment of SpondyloArthritis International Society (ASAS) classification criteria for axial spondyloarthritis (all participant demographics provided in Supplementary Table 2)32. Healthy controls were recruited through the Volunteers for Health registry at Washington University in St. Louis. All were free of autoimmune diseases. HLA-B*27 positivity was confirmed through PCR in the laboratory.
Biospecimen collection
Peripheral blood samples were obtained by venipuncture and collected into EDTA tubes. Peripheral blood mononuclear cells (PBMCs) were isolated by Ficoll-Hypaque density gradient centrifugation. PBMCs were cryopreserved in FBS containing 10% dimethyl sulfoxide (DMSO) and stored at −140 °C until analysis26.
Flow cytometry
Frozen PBMCs were thawed and washed twice with RPMI + 10% FBS. A total of 10 million cells were used for flow cytometric assessment of HLA-B*27-restricted CD8+ T cells specific to PSG5 and NP.383-391 (SRYWAIRTR). PSG5 tetramers were conjugated using streptavidin–PE (Agilent, PJRS25-1) and used at a 1:50 dilution. NP.383-391 tetramers were conjugated using streptavidin–BV650 (BioLegend, 405232) and used at a 1:100 dilution24. Cells were washed with PBS, stained with fixable viability dye (Thermo Fisher Scientific) at 4 °C for 20 min and then washed once with FACS buffer (PBS containing 1% FBS). Cells were resuspended in Fc block (Thermo Fischer Scientific) for 10 min at room temperature. They were then incubated with tetramers and surface antibodies (all antibody reagents provided in Supplementary Table 3) for 30 min at room temperature. Cells were washed twice with FACS buffer and fixed with 2% PFA for flow cytometry. Data were collected on Cytek Aurora flow cytometer (Cytek Biosciences) and analyzed with FlowJo (version 10; Tree Star).
Expression of AS/AAU TCR in Expi293 mammalian expression system
TCR α-chains and β-chains were cloned into the pD649 mammalian expression vector. The α-chain construct included a C-terminal basic leucine zipper, while the β-chain construct contained a C-terminal acidic leucine zipper and a BirA biotinylation tag. Both chains were cotransfected into Expi293 cells (Thermo Fisher, A14527) following the manufacturer’s protocol33.
After a 4-day expression period, cultures were centrifuged at 2,000g for 10 min to collect the supernatant. The supernatant was mixed with an equal volume of PBS (pH 7.2, Gibco) and incubated with 2 ml of Ni-NTA agarose slurry (Qiagen) per 50 ml of supernatant at 4 °C overnight with gentle rotation. The mixture was passed through a gravity column to isolate resin-bound protein. The resin was washed with 30 column volumes (CVs) of PBS containing 20 mM imidazole and eluted with 10 CVs of PBS containing 200 mM imidazole.
The eluted protein was concentrated using a Millipore filter (30-kDa molecular weight cutoff) to a final volume of ~500 µl and subsequently biotinylated overnight using BirA. Final purification was performed by size-exclusion chromatography using a Superdex 200 Increase column (GE Healthcare) equilibrated in PBS pH 7.2.
Fractions were analyzed for heterodimeric TCR formation and successful biotinylation by SDS–PAGE (under reducing and nonreducing conditions) and streptavidin shift assays. Peak fractions were pooled, quantified by absorbance at 280 nm, aliquoted and stored at −80 °C until use.
Construction and selection of yeast-displayed peptide–HLA-B*27:05 libraries
The peptide–HLA-B27:05 library was displayed on yeast as a single-chain trimer (SCT) comprising a 9-mer peptide, β2-microglobulin (β2m) and the HLA-B27:05 heavy chain, linked by flexible linkers. To enhance stability, the construct included previously reported substitutions (Y84A, L5M, H114Y and A153D with C67S to prevent aberrant disulfide formation) and was assembled following established protocols24.
Peptide libraries were generated using primers encoding predefined codons (all primers are provided in Supplementary Table 4). To ensure optimal binding to HLA-B*27:05 and specificity toward AS-associated TCRs, anchor residues at positions P2 and P8 were fixed as arginine and proline, respectively. The remaining positions were encoded using NNK degenerate codons to introduce diversity. Each 9-mer library was constructed independently and fused to a C-terminal MYC epitope tag for flow cytometry detection. Library complexity was estimated by colony counting after yeast electroporation.
Transformed yeast was grown in SDCAA medium (pH 4.5), induced in SGCAA medium (pH 4.5) and subjected to iterative selection. Before positive selection, negative depletion was performed by incubating yeast (10× library diversity) with streptavidin-coated magnetic beads at 4 °C for 1 h to remove nonspecific binders. Cells were passed through LS columns (Miltenyi) under a magnetic field and washed three times; the flowthrough was collected for subsequent enrichment.
For positive selection, yeast were incubated with 400 nM biotinylated soluble TCR preconjugated to streptavidin-coated MACS beads at 4 °C for 3 h. Bound cells were retained on LS columns, eluted and expanded overnight in SDCAA medium. This induction and selection cycle was repeated for a total of four rounds. Selected yeast populations were stored at 4 °C and used for next-generation sequencing analysis.
In addition, TCRs 4.2, 4.3 and 4.4 were analyzed using previously published peptide–HLA-B*27:05 library data, in which P2 and P9 were fixed while P8 was unconstrained24.
Deep sequencing of pHLA libraries
Yeast DNA was extracted from approximately 1 × 107 yeast cells per selection round using the Zymoprep II Miniprep Kit (Zymo Research). Peptide-encoding regions were PCR-amplified over 30 cycles using primers containing Illumina sequencing adaptors, unique 8-nt unique molecular identifiers and round-specific barcodes to distinguish between selection rounds (Supplementary Table 4).
PCR products were purified by agarose gel extraction and quantified using the Qubit dsDNA HS assay kit (Thermo Fisher Scientific). Libraries were sequenced on an Illumina MiSeq platform using a 2× 150-bp v2 kit. Forward reads were demultiplexed in Geneious Prime 2024.0.7 on the basis of barcode sequences to separate individual selection rounds. Reads containing frameshifts or premature stop codons were removed from further analysis.
Identical peptide sequences were counted and clustered using custom Python scripts and Unix shell commands. For each TCR, peptide counts from selection rounds 1–4 were aggregated into a structured dataset, which served as input for downstream machine learning-based prediction and analysis.
Deep sequencing binding binarization and dataset preparation
Peptide counts from selection rounds 1–4 were used to binarize binding labels. Peptides were designated as positive binders if they had a count ≥ 5 in round 4. Negative binders were defined as peptides with a count of zero across rounds 2–4.
T cell activation in coculture assays
TCRs were cloned into the pHR lentiviral vector. Full-length α-chain and β-chain constructs were cloned separately and cotransfected with packaging plasmids pMD2.G and pSPAX2 into HEK293T cells (American Type Culture Collection (ATCC), CRL-3216) using Fugene (Promega) in DMEM supplemented with 10% FBS. Lentiviral supernatants were isolated 60 h after transfection, clarified by centrifugation and used to transduce 1.5 × 106 CD8+ SKW-3 cells by spinfection in the presence of 8 µg ml−1 polybrene (Millipore Sigma). Transduced cells were cultured in RPMI complete medium and expanded for 5–7 days. TCR surface expression was validated by flow cytometry using PE anti-CD3 (UCHT1, BioLegend, 300408) or PE anti-TCR (IP26, BioLegend, 984702) antibodies.
Similarly, full-length HLA-B*27:05 and β2m were cloned into the pHR vector and packaged in HEK293T cells under identical conditions. Lentivirus was collected 60 h after transfection and used to transduce 1.5 × 106 K-562 cells (ATCC, CCL-243) following the same spinfection protocol. After expansion for 5–7 days, surface HLA expression was assessed by staining with APC anti-HLA-A/B/C antibody (W6/32, BioLegend, 311410) and analyzed by flow cytometry.
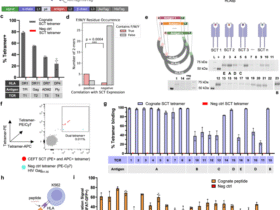
For antigen presentation, HLA-B*27:05-transduced K-562 cells were labeled with CellTrace violet (Invitrogen) for 20 min and incubated with 15 predicted cross-reactive peptides at 37 °C in 5% CO2 for 1–2 h. Peptides (≥70% purity, Elim Biopharm) were dissolved in DMSO at 10 mM and 1 µl of peptide stock was added to 100 µl of K-562 cells to reach a final concentration of 100 µM. After peptide loading, cells were washed with RPMI and cocultured with TCR-transduced SKW-3 cells at a 1:1 ratio for 14–18 h.
T cell activation was assessed by staining for APC anti-CD69 (FN50, BioLegend, 310910) and PE anti-CD3 (OKT3 or UCHT1, BioLegend, 317308 or 300408), followed by flow cytometric analysis (CytoFLEX, Beckman Coulter) and analysis using GraphPad Prism (versions 8.4 and 9.0).
T cell activation data binarization
To normalize T cell activation data, the CD69% of CD3+ cells stimulated with a null control peptide was subtracted from the CD69% of CD3+ cells stimulated with each test peptide. This normalization was performed independently for each replicate, followed by averaging across replicates. For binarization, peptides eliciting >3% CD69 expression were classified as activators.
Expression of HLA-B*27 as inclusion bodies in Escherichia
coli
The HLA-B*27:05 heavy chain and human β2m were expressed in E. coli as inclusion bodies. The respective genes were cloned into the pET26b vector and transformed into E. coli BL21(DE3) competent cells. A single colony was picked and cultured overnight in 10 ml of LB medium supplemented with 50 µg ml−1 kanamycin at 37 °C with shaking at 250 rpm for 12–16 h. The overnight culture was then inoculated into 1 L of LB medium containing 50 µg ml−1 kanamycin and incubated at 37 °C, 300 rpm until the optical density at 600 nm (OD600) reached 0.5–0.7.
Protein expression was induced by adding IPTG to a final concentration of 1 mM, followed by continued shaking at 37 °C for an additional 3 h. Bacterial cultures were harvested by centrifugation at 6,000g for 15 min at 4 °C. Pelleted cells were resuspended in 50 ml of lysis buffer (50 mM Tris-HCl pH 8.0, 100 mM NaCl, 10 mM DTT, 1% Triton X-100, 5 mM MgCl2 and 0.2 mM PMSF) and lysed by sonication using a 2 min on–off cycle, repeated three times.
The lysate was centrifuged at 10,000g for 15 min at 4 °C. Subsequently, the pellet was resuspended in 50 ml of wash buffer (50 mM Tris-HCl pH 8.0, 100 mM NaCl, 0.5% Triton X-100, 1 mM EDTA, 1 mM DTT and 0.2 mM PMSF) and subjected to the same sonication cycle (2 min on, 2 min off, repeated four times), followed by centrifugation at 10,000g for 15 min. This wash step was repeated once more with wash buffer.
The inclusion body pellet was solubilized in 25 ml of solubilization buffer containing 8 M urea, 20 mM Tris-HCl pH 8.0, 0.5 mM EDTA and 1 mM DTT. Insoluble debris was removed by centrifugation at 60,000g for 30 min at 4 °C and the clarified supernatant was collected. Solubilized protein was aliquoted and stored at −80 °C until use in refolding experiments.
Refolding of pMHC
The refolding buffer was prepared with 100 mM Tris-HCl pH 8.0, 400 mM arginine, 0.5 mM oxidized glutathione, 5 mM reduced glutathione and 2 mM EDTA. Peptide (20 mg) was dissolved in DMSO and added to 1 L of refolding buffer.
For each liter of refolding buffer, 20 mg of HLA-B*27:05 heavy chain inclusion body and 20 mg of human β2m inclusion body were premixed and added dropwise into the refolding buffer under gentle stirring. The refolding mixture was transferred into dialysis tubing (Spectrum Labs) and dialyzed against 10 L of 20 mM Tris-HCl pH 8.0, with buffer changes every 12 h for a total of four exchanges.
Following dialysis, refolded pMHC complexes were filtered through a 0.22-µm membrane, concentrated to ~1 ml and subjected to ion-exchange chromatography using a Mono Q column (GE Healthcare). Final quality assessment was performed by size-exclusion chromatography on a Superdex 200 Increase column (GE Healthcare) using an ÄKTA purifier system.
Crystallization of AS TCR–pHLA-B*27:05 complexes
AS TCRs expressed in GnTI− Expi293 cells (Thermo Fisher, A39240) were treated with Endo H, 3C protease and carboxypeptidases A and B at 4 °C for 24 h to remove N-linked glycans and C-terminal leucine zipper tags. The reaction mixture of TCR and refolded pMHC was subsequently purified by size-exclusion chromatography using a Superdex 200 column (GE Healthcare). Protein complexes were crystallized by sitting-drop vapor diffusion: AS19.2–YEIH–HLA-B*27:05 was at 10 mg ml−1 using 0.2 M sodium malonate pH 6.4, 17% PEG 3350 and AS19.2–PSG5–HLA-B*27:05 at 17 mg ml−1 with 0.2 M sodium formate and 20% PEG 3350. Crystals were cryoprotected with addition of 30% glycerol and flash-cooled in liquid nitrogen for data collection.
Structure determination and refinement
Diffraction data were collected at Stanford Synchrotron Radiation Lightsource (SSRL) beamline 12-2. Intensities were processed using XDS34. The YEIH structure was determined by molecular replacement using the PHASER (version 2.5.5) program with components of PDB 7N2R as the search models and the PSG5 structure used the rebuilt TCR 19.2 chains from the YEIH structure with HLA chains from PDB 7N2R. Model building was performed manually in Coot (version 0.9.6), and structure refinement was conducted using the PHENIX (version 2.5.5) software suite35,36. Simulated annealing omit electron density maps are presented in Supplementary Fig. 8. Structural analyses were carried out with programs from the CCP4 package (version 7.0)37. Crystallographic software was installed and maintained by SBGrid38. Data collection and refinement statistics are provided in Supplementary Table 5. Final coordinates and structure factors were deposited to the RCSB Protein Data Bank under accession codes 9PBG and 9PBH. Diffraction images were deposited to the SBGrid data bank under accession numbers 1178 for YEIH and 1179 for PSG5.
Motif analysis and sequence logos
Positive binder peptides for each TCR were used to generate motif using MoDec (version 1.2)39 where the best motif was selected across five runs allowing for a single motif with the background amino acid frequencies drawn from the human proteome.
JS distances were calculated using Scipy (scipy.spatial.distance.jensenshannon) by taking the sum of the distances for all putative TCR contact residues (positions 0, 2, 3, 5 and 6). MDS was then performed using sklearn (sklearn.manifold.MDS) with n_components = 2 using the computed JS distances.
Machine learning architecture
Our model architecture is centered on a single pretrained encoder ESM2 (650-million-parameter model), to encode and extract representations for both the CDR3 α/β and antigen peptide sequence to obtain sequence-wise representations. Each sequence was independently embedded using mean-pooled position-wise representations. Additionally, the position-wise embeddings concatenated with one-hot encoded sequence was further processed and passed through a six-layer transformer encoder. The output of the transformer encoder and mean-pooled position-wise representations were concatenated to form the final embeddings for each peptide and CDR3 α/β. For our β-chain models, a joint representation was then computed as the outer product between the CDR3 and peptide for the sequence-wise embeddings and position-wise representations (two channels). For our α/β-chain models, the outer product was computed for each CDR3 and peptide and concatenated in the channel dimension (four channels). Joint embeddings were then processed through a three-layer convolutional neural network, followed by a single-layer multilayer perceptron to predict binding probability. Binary cross-entropy loss was used for optimization. All model parameters, including the ESM2 encoder, were fine-tuned during training.
Pretraining on VDJdb
Before training on deep sequencing data, we pretrained the model on VDJdb. We extracted CDR3α/β–peptide pairs belonging to MHC class I. We used the cognate pairs provided as positives. Negative pairs were generated by uniformly sampling noncognate peptides from the VDJdb pool each epoch. An equal number of positive and negative examples were sampled each epoch. The data were split into training (95%) and validation (5%) sets. The model was trained using the Adam optimizer (learning rate: 1 × 10−6) until convergence, with early stopping based on a validation patience of ten epochs.
Training single-TCR models
Fine-tuning was performed on each binarized TCR dataset using the pretrained VDJdb model. Each dataset was split into training (70%), validation (15%) and test (15%) sets. Training used the same hyperparameters as during pretraining. To address class imbalance, where negative samples greatly outnumber positives, we applied a weighted binary cross-entropy loss, with class weights set inversely proportional to the number of positive and negative examples in the training set. This ensured that each class contributed equally. Performance was evaluated on the held-out test set using the average precision (AP) and AUROC. For TCR 19.2, we additionally evaluated the model on its five mutant variants (C1–C5) to assess generalization to mutant CDR3β variants.
Training joint and LOO TCR models
We fine-tuned the VDJdb-pretrained model on a concatenated dataset comprising all 16 individual TCR datasets. The same 70:15:15 train–validation–test split was used to enable fair comparison to individual TCR models. For the joint 19.2 model (Fig. 5d), we combined the WT and five mutant (C1–C5) datasets in training. To assess generalizability, we trained 16 LOO models, each using data from 15 TCRs and evaluating on the held-out TCR. As TCRs 28.1 and 135.7 share the same CDR3β, both were excluded whenever either was held out to prevent data leakage. Consequently, although 16 LOO models were trained, only 15 were unique. Training hyperparameters were identical to those used in prior experiments.
Generating human proteome 9-mer peptide library and nominating peptides for activation study
We used NetMHCpan-4.1 and the UniProt human reference proteome (UP000005640) to predict all 9-mer peptides with HLA-B*27:05-binding potential. Peptides with percentage rank < 2 (weak binders) and percentage rank < 0.5 (strong binders) were subselected, yielding 293,467 unique peptides. On the basis of prior knowledge that the tested TCRs exhibit a strong preference for proline at P8 of the peptide, we further filtered the dataset to include only peptides containing proline at P8, yielding 12,357 unique peptides. To nominate peptides for functional testing, we applied each single-TCR model to score this peptide library. For each TCR, peptides with predicted binding scores > 0.95 were selected. We then subselected peptides predicted to bind at least five TCRs. Ranking each of the subselected peptides by the number of TCRs predicted to bind, the top 15 peptides were selected. The maximum number of TCRs that a single peptide was predicted to bind was 12. For model performance evaluation and benchmarking against other models (AUROC and AP), we excluded TCRs 4.1, 4.4 and 135.7, as none of the selected peptides activated these TCRs. To benchmark against peptide to MHC affinity-based prediction, we selected the top 20 peptides from the human proteome with the highest predicted affinity for HLA-B*27:05 using NetMHCpan-4.1. These peptides were experimentally tested for T cell activation to serve as high-affinity nonactivating controls.
Calculating TCR distances
To assess relatedness between TCRs and TCR neighborhoods in the LOO setting, we computed several distance metrics. The JS distance was computed position-wise between amino acid distributions derived from peptide enrichment profiles obtained through yeast-display screening and averaged across non-excluded peptide positions. For each included position i, amino acid distributions pi and qi, representing the peptide-binding motifs of two TCRs being compared, were compared using the following definitions:
$$\begin{array}{rcl}d_{\mathrm{JS}}\left(p,q\right)&=&\frac{1}{\left|I\right|}\mathop{\sum }\limits_{i\in I}\sqrt{\frac{D_{\mathrm{KL}}\left(p_i\parallel m_i\right)+D_{\mathrm{KL}}\left(q_i\parallel m_i\right)}{2}},\quad m_i=\frac{1}{2}\left(p_i+q_i\right),\\ D_{\mathrm{KL}}\left(p_i\parallel m_i\right)&=&\mathop{\sum }\limits_{a\in\mathcal{A}}p_i\left(a\right)\log\frac{p_i\left(a\right)}{m_i\left(a\right)},\quad I=\left\{1,\ldots,L\right\}\setminus\left\{2,8,9\right\}\end{array}$$
Here, DKL denotes the Kullback-Leibler divergence, \(\mathcal{A}\) denotes the amino acid alphabet and I denotes the set of included peptide positions. Positions P2, P8 and P9 were excluded from distance calculations because P2 corresponds to a canonical MHC anchor position, while P8 and P9 were fixed in subsets of peptide libraries during library construction, resulting in limited amino acid variability at those positions among positive binders. This motif-based JS distance was used as the ground-truth measure of similarity between TCRs. Edit distance was computed using CDR3β amino acid sequences only. TCRdist3 was computed using TCRβ-chain information only, including the CDR3β sequence together with the corresponding V- and J-gene annotations, to ensure a fair comparison with our TCRβ-centric model. Mahalanobis distance was calculated as
$$d_{\mathrm{Mah}}\left(x,\mu\right)=\sqrt{\left(x-\mu\right)^T S^{-1}\left(x-\mu\right)}$$
where x denotes the sequence-level representations of the held-out CDR3β sequence after the ESM module, μ denotes the mean vector of training-set sequence-level representations after the ESM module and S−1 denotes the Moore-Penrose pseudoinverse covariance matrix computed from the training-set sequence-level representations.
Because TCRs 28.1 and 135.7 share identical CDR3β sequences, each TCR was excluded from the other’s distance calculations to maintain consistency with the LOO training procedure.
AlphaFold3 and tFold-TCR predictions
To benchmark against existing complex structure-prediction algorithms, we evaluated both AlphaFold3 (a general multimolecular structure predictor) and tFold-TCR (an algorithm specifically trained for TCR–pMHC complex modeling). For each peptide in our activation study, we supplied the full sequences for each component of the TCR–pMHC complex as input to each method using default inference settings. Both AlphaFold3 and tFold generate interface confidence metrics intended to reflect the reliability of predicted intermolecular contacts. We extracted the ipTM score from each model and used this score as the structure-based predictor of TCR activation.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.















Leave a Reply